Target Information
| Target General Information | Top | |||||
|---|---|---|---|---|---|---|
| Target ID |
T15510
(Former ID: TTDI02453)
|
|||||
| Target Name |
Insulin receptor substrate-1 (IRS1)
|
|||||
| Synonyms |
Insulin receptor substrate 1; IRS-1
Click to Show/Hide
|
|||||
| Gene Name |
IRS1
|
|||||
| Target Type |
Clinical trial target
|
[1] | ||||
| Disease | [+] 1 Target-related Diseases | + | ||||
| 1 | Solid tumour/cancer [ICD-11: 2A00-2F9Z] | |||||
| Function |
When phosphorylated by the insulin receptor binds specifically to various cellular proteins containing SH2 domains such as phosphatidylinositol 3-kinase p85 subunit or GRB2. Activates phosphatidylinositol 3-kinase when bound to the regulatory p85 subunit. May mediate the control of various cellular processes by insulin.
Click to Show/Hide
|
|||||
| UniProt ID | ||||||
| Sequence |
MASPPESDGFSDVRKVGYLRKPKSMHKRFFVLRAASEAGGPARLEYYENEKKWRHKSSAP
KRSIPLESCFNINKRADSKNKHLVALYTRDEHFAIAADSEAEQDSWYQALLQLHNRAKGH HDGAAALGAGGGGGSCSGSSGLGEAGEDLSYGDVPPGPAFKEVWQVILKPKGLGQTKNLI GIYRLCLTSKTISFVKLNSEAAAVVLQLMNIRRCGHSENFFFIEVGRSAVTGPGEFWMQV DDSVVAQNMHETILEAMRAMSDEFRPRSKSQSSSNCSNPISVPLRRHHLNNPPPSQVGLT RRSRTESITATSPASMVGGKPGSFRVRASSDGEGTMSRPASVDGSPVSPSTNRTHAHRHR GSARLHPPLNHSRSIPMPASRCSPSATSPVSLSSSSTSGHGSTSDCLFPRRSSASVSGSP SDGGFISSDEYGSSPCDFRSSFRSVTPDSLGHTPPARGEEELSNYICMGGKGPSTLTAPN GHYILSRGGNGHRCTPGTGLGTSPALAGDEAASAADLDNRFRKRTHSAGTSPTITHQKTP SQSSVASIEEYTEMMPAYPPGGGSGGRLPGHRHSAFVPTRSYPEEGLEMHPLERRGGHHR PDSSTLHTDDGYMPMSPGVAPVPSGRKGSGDYMPMSPKSVSAPQQIINPIRRHPQRVDPN GYMMMSPSGGCSPDIGGGPSSSSSSSNAVPSGTSYGKLWTNGVGGHHSHVLPHPKPPVES SGGKLLPCTGDYMNMSPVGDSNTSSPSDCYYGPEDPQHKPVLSYYSLPRSFKHTQRPGEP EEGARHQHLRLSTSSGRLLYAATADDSSSSTSSDSLGGGYCGARLEPSLPHPHHQVLQPH LPRKVDTAAQTNSRLARPTRLSLGDPKASTLPRAREQQQQQQPLLHPPEPKSPGEYVNIE FGSDQSGYLSGPVAFHSSPSVRCPSQLQPAPREEETGTEEYMKMDLGPGRRAAWQESTGV EMGRLGPAPPGAASICRPTRAVPSSRGDYMTMQMSCPRQSYVDTSPAAPVSYADMRTGIA AEEVSLPRATMAAASSSSAASASPTGPQGAAELAAHSSLLGGPQGPGGMSAFTRVNLSPN RNQSAKVIRADPQGCRRRHSSETFSSTPSATRVGNTVPFGAGAAVGGGGGSSSSSEDVKR HSSASFENVWLRPGELGGAPKEPAKLCGAAGGLENGLNYIDLDLVKDFKQCPQECTPEPQ PPPPPPPHQPLGSGESSSTRRSSEDLSAYASISFQKQPEDRQ Click to Show/Hide
|
|||||
| 3D Structure | Click to Show 3D Structure of This Target | AlphaFold | ||||
| ADReCS ID | BADD_A06414 | |||||
| HIT2.0 ID | T28K6X | |||||
| Cell-based Target Expression Variations | Top | |||||
|---|---|---|---|---|---|---|
| Cell-based Target Expression Variations | ||||||
| Drug Binding Sites of Target | Top | |||||
|---|---|---|---|---|---|---|
| Ligand Name: L-betagamma-meATP | Ligand Info | |||||
| Structure Description | Structure of the Insulin-like Growth Factor 1 Receptor Kinase | PDB:1K3A | ||||
| Method | X-ray diffraction | Resolution | 2.10 Å | Mutation | Yes | [3] |
| PDB Sequence |
GEYVNIEF
|
|||||
|
|
||||||
| Click to View More Binding Site Information of This Target with Different Ligands | ||||||
| Different Human System Profiles of Target | Top |
|---|---|
|
Human Similarity Proteins
of target is determined by comparing the sequence similarity of all human proteins with the target based on BLAST. The similarity proteins for a target are defined as the proteins with E-value < 0.005 and outside the protein families of the target.
A target that has fewer human similarity proteins outside its family is commonly regarded to possess a greater capacity to avoid undesired interactions and thus increase the possibility of finding successful drugs
(Brief Bioinform, 21: 649-662, 2020).
Human Tissue Distribution
of target is determined from a proteomics study that quantified more than 12,000 genes across 32 normal human tissues. Tissue Specificity (TS) score was used to define the enrichment of target across tissues.
The distribution of targets among different tissues or organs need to be taken into consideration when assessing the target druggability, as it is generally accepted that the wider the target distribution, the greater the concern over potential adverse effects
(Nat Rev Drug Discov, 20: 64-81, 2021).
Human Pathway Affiliation
of target is determined by the life-essential pathways provided on KEGG database. The target-affiliated pathways were defined based on the following two criteria (a) the pathways of the studied target should be life-essential for both healthy individuals and patients, and (b) the studied target should occupy an upstream position in the pathways and therefore had the ability to regulate biological function.
Targets involved in a fewer pathways have greater likelihood to be successfully developed, while those associated with more human pathways increase the chance of undesirable interferences with other human processes
(Pharmacol Rev, 58: 259-279, 2006).
Biological Network Descriptors
of target is determined based on a human protein-protein interactions (PPI) network consisting of 9,309 proteins and 52,713 PPIs, which were with a high confidence score of ≥ 0.95 collected from STRING database.
The network properties of targets based on protein-protein interactions (PPIs) have been widely adopted for the assessment of target’s druggability. Proteins with high node degree tend to have a high impact on network function through multiple interactions, while proteins with high betweenness centrality are regarded to be central for communication in interaction networks and regulate the flow of signaling information
(Front Pharmacol, 9, 1245, 2018;
Curr Opin Struct Biol. 44:134-142, 2017).
Human Similarity Proteins
Human Tissue Distribution
Human Pathway Affiliation
Biological Network Descriptors
|
|
|
There is no similarity protein (E value < 0.005) for this target
|
|
Note:
If a protein has TS (tissue specficity) scores at least in one tissue >= 2.5, this protein is called tissue-enriched (including tissue-enriched-but-not-specific and tissue-specific). In the plots, the vertical lines are at thresholds 2.5 and 4.
|
| KEGG Pathway | Pathway ID | Affiliated Target | Pathway Map |
|---|---|---|---|
| cGMP-PKG signaling pathway | hsa04022 | Affiliated Target |

|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| FoxO signaling pathway | hsa04068 | Affiliated Target |

|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Autophagy - animal | hsa04140 | Affiliated Target |
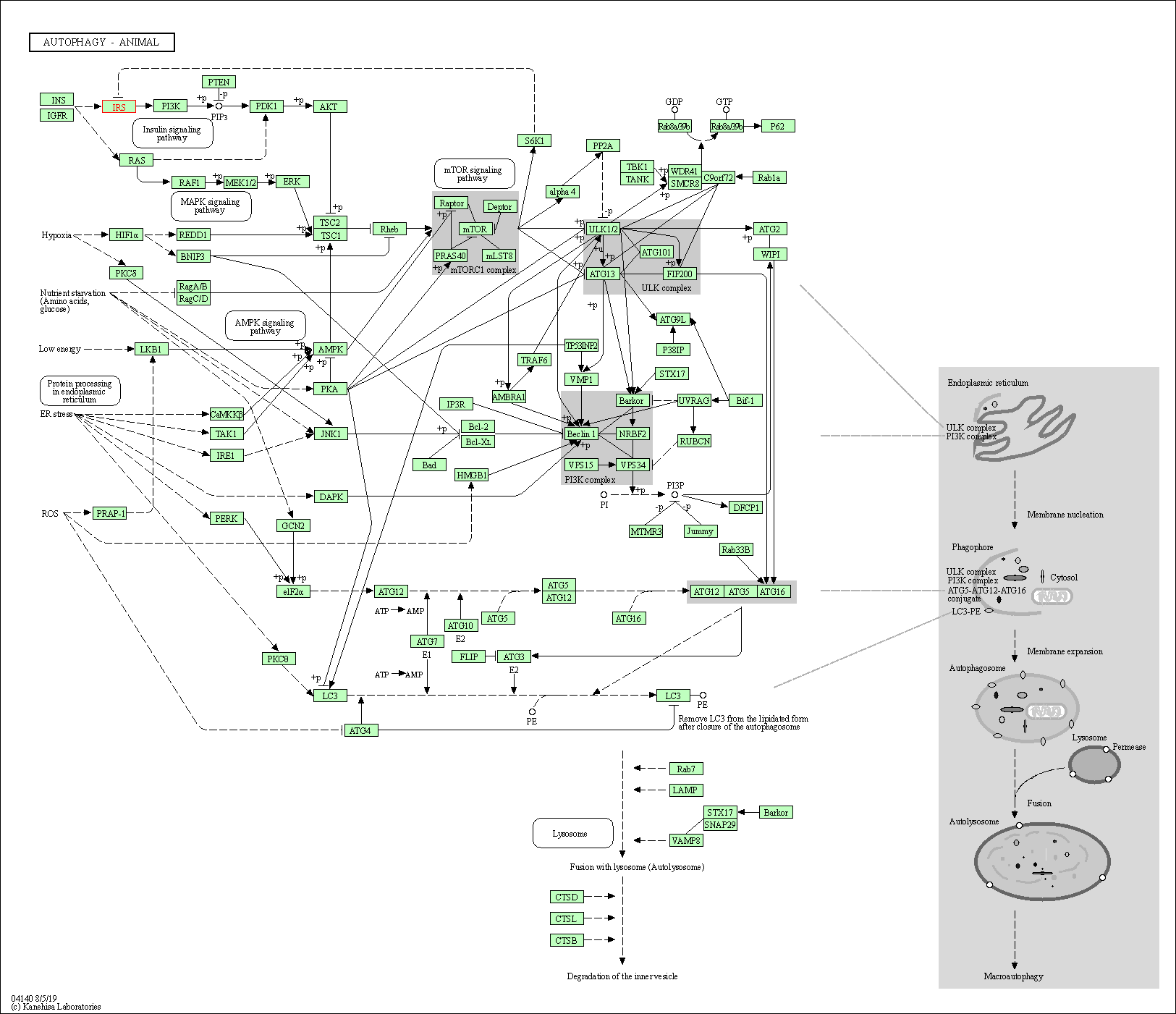
|
| Class: Cellular Processes => Transport and catabolism | Pathway Hierarchy | ||
| mTOR signaling pathway | hsa04150 | Affiliated Target |

|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| PI3K-Akt signaling pathway | hsa04151 | Affiliated Target |

|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| AMPK signaling pathway | hsa04152 | Affiliated Target |

|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Longevity regulating pathway | hsa04211 | Affiliated Target |
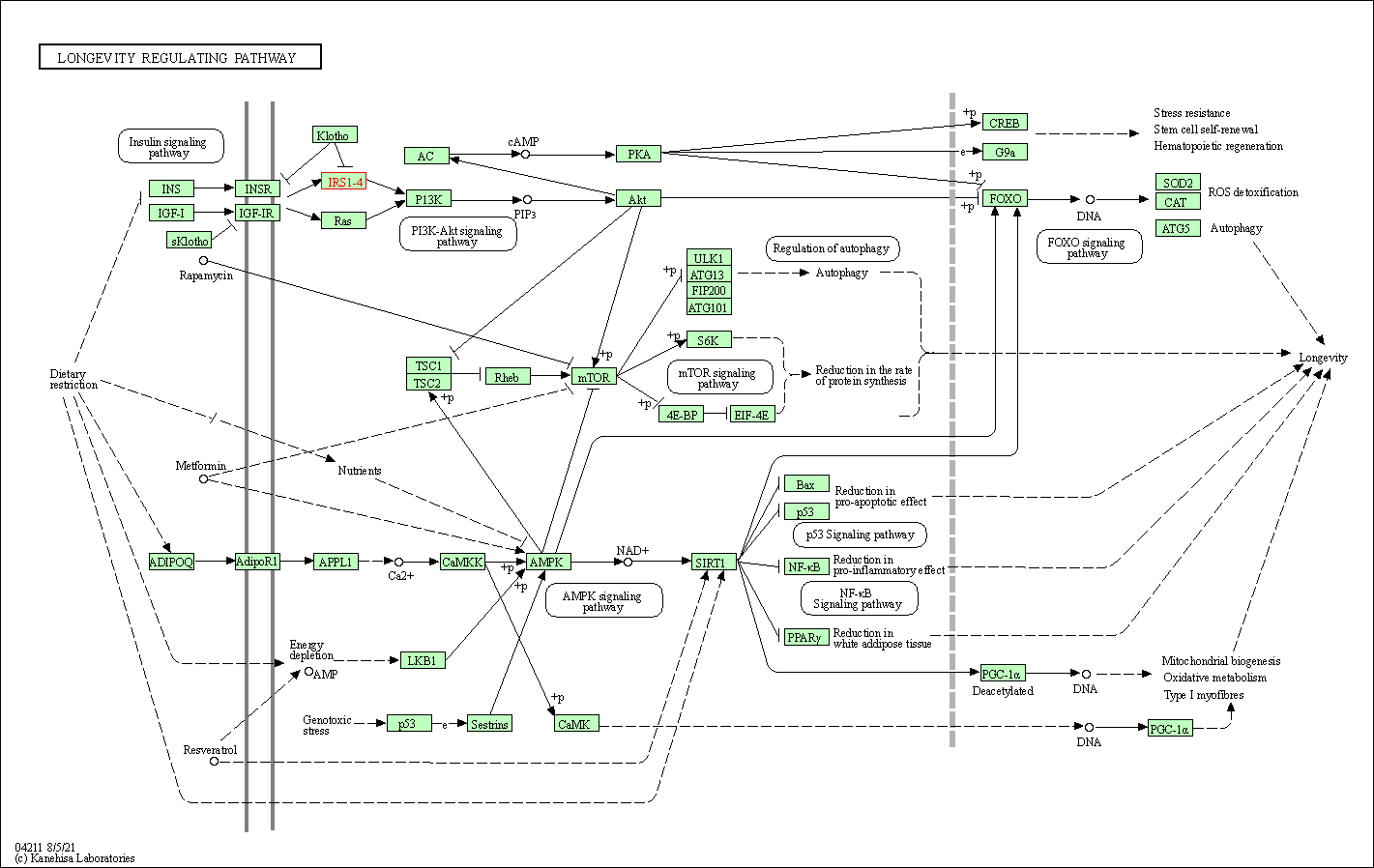
|
| Class: Organismal Systems => Aging | Pathway Hierarchy | ||
| Longevity regulating pathway - multiple species | hsa04213 | Affiliated Target |

|
| Class: Organismal Systems => Aging | Pathway Hierarchy | ||
| Neurotrophin signaling pathway | hsa04722 | Affiliated Target |
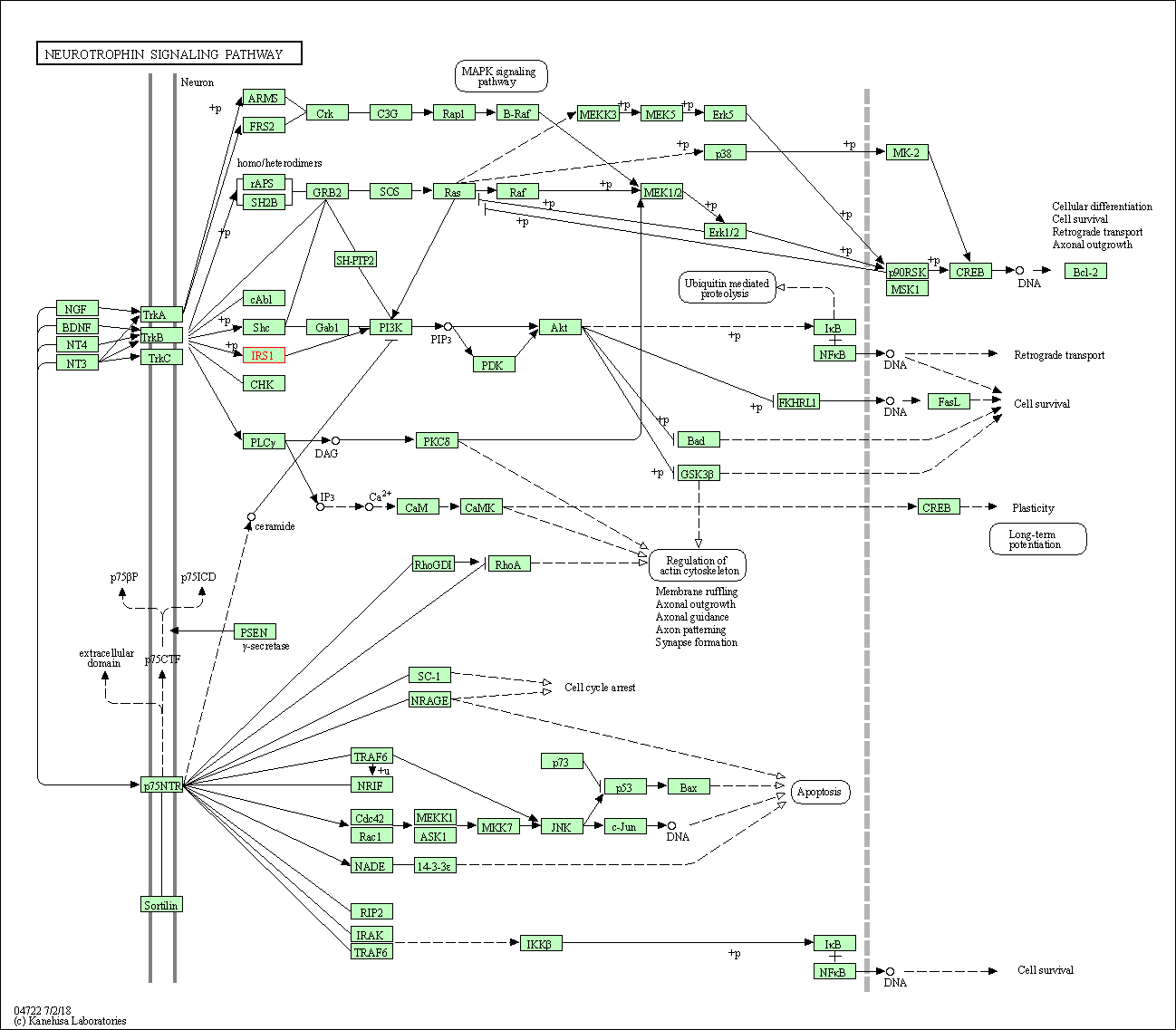
|
| Class: Organismal Systems => Nervous system | Pathway Hierarchy | ||
| Insulin signaling pathway | hsa04910 | Affiliated Target |
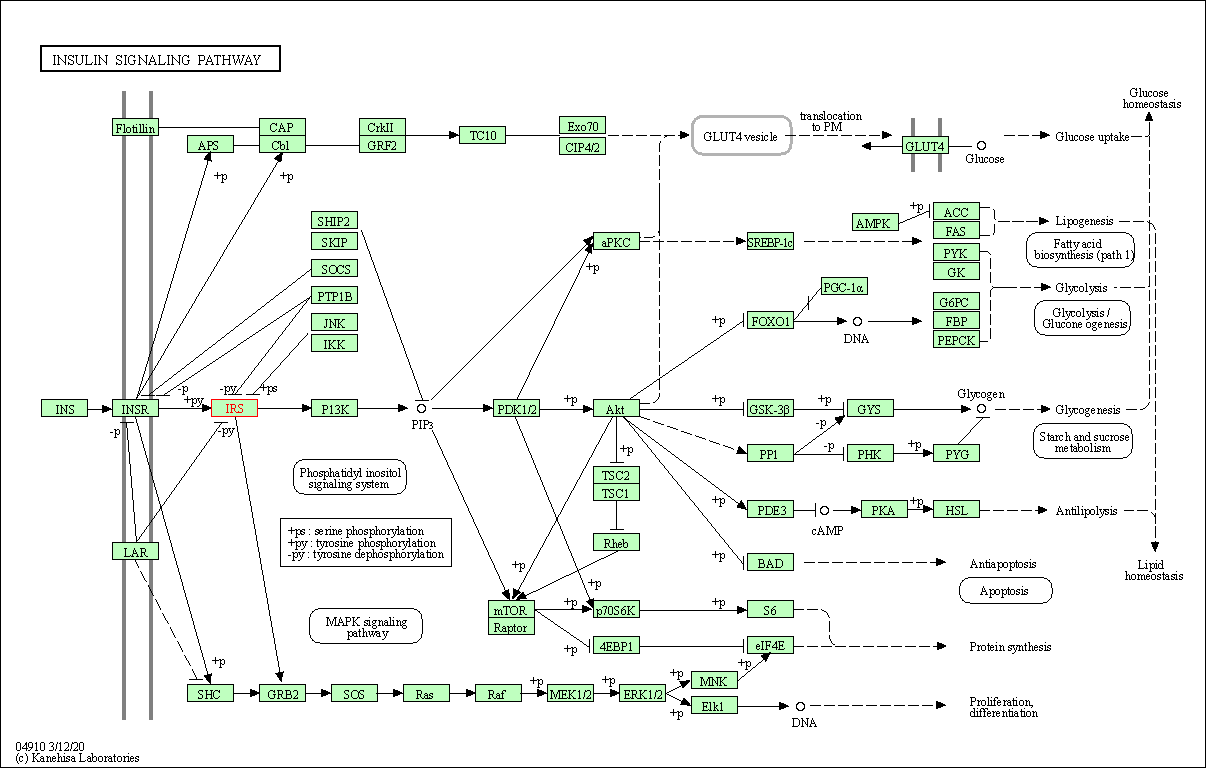
|
| Class: Organismal Systems => Endocrine system | Pathway Hierarchy | ||
| Adipocytokine signaling pathway | hsa04920 | Affiliated Target |
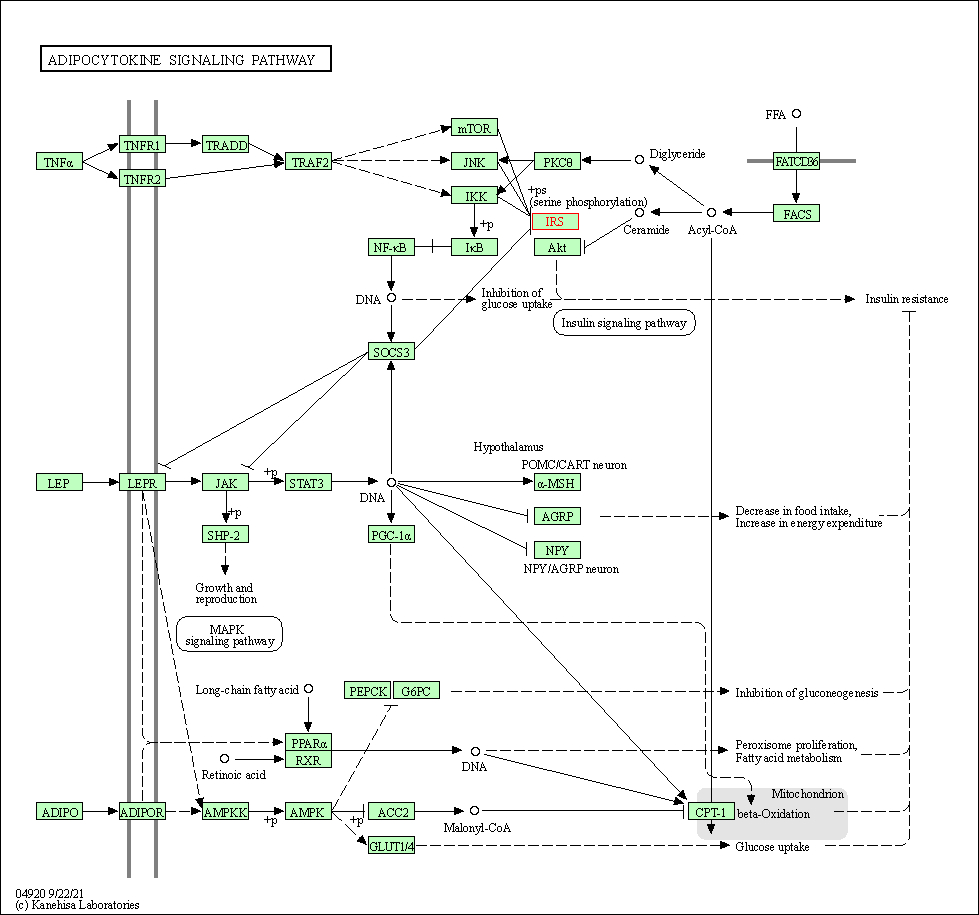
|
| Class: Organismal Systems => Endocrine system | Pathway Hierarchy | ||
| Regulation of lipolysis in adipocytes | hsa04923 | Affiliated Target |
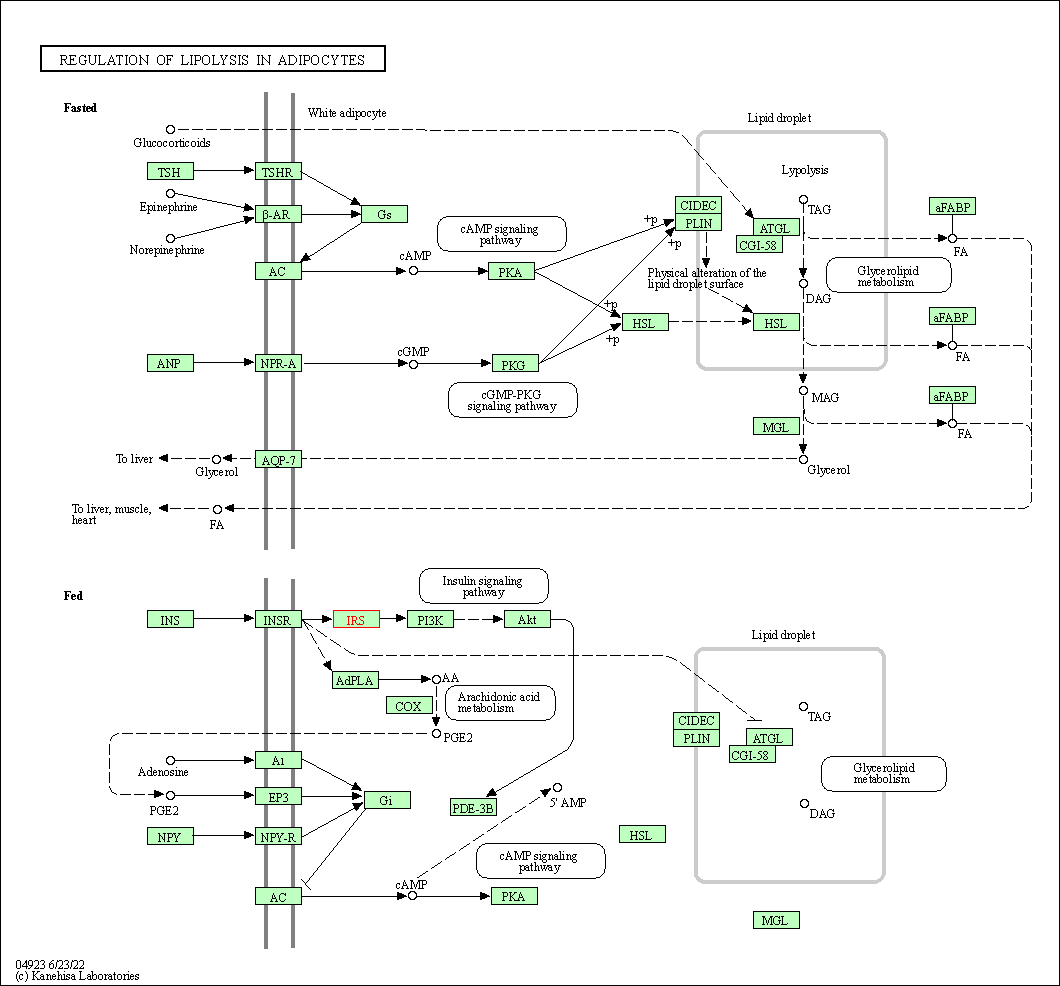
|
| Class: Organismal Systems => Endocrine system | Pathway Hierarchy | ||
| Growth hormone synthesis, secretion and action | hsa04935 | Affiliated Target |
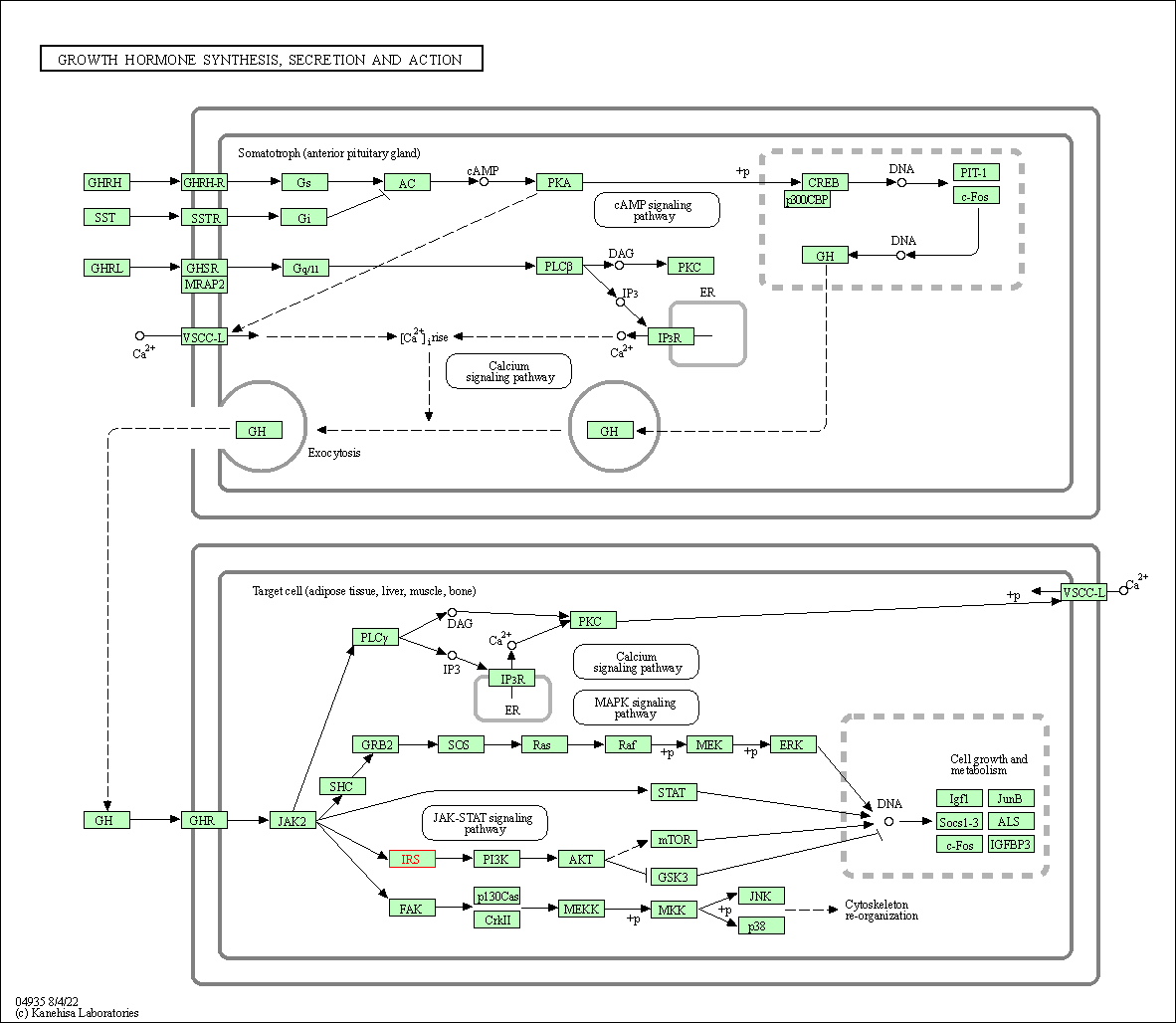
|
| Class: Organismal Systems => Endocrine system | Pathway Hierarchy | ||
| Aldosterone-regulated sodium reabsorption | hsa04960 | Affiliated Target |

|
| Class: Organismal Systems => Excretory system | Pathway Hierarchy | ||
| Click to Show/Hide the Information of Affiliated Human Pathways | |||
| Degree | 45 | Degree centrality | 4.83E-03 | Betweenness centrality | 1.39E-03 |
|---|---|---|---|---|---|
| Closeness centrality | 2.64E-01 | Radiality | 1.45E+01 | Clustering coefficient | 2.47E-01 |
| Neighborhood connectivity | 5.84E+01 | Topological coefficient | 5.66E-02 | Eccentricity | 11 |
| Download | Click to Download the Full PPI Network of This Target | ||||
| Target Regulators | Top | |||||
|---|---|---|---|---|---|---|
| Target-regulating microRNAs | ||||||
| Target-interacting Proteins | ||||||
| References | Top | |||||
|---|---|---|---|---|---|---|
| REF 1 | MicroRNA-145 Negatively Regulates Cell Proliferation Through Targeting IRS1 in Isolated Ovarian Granulosa Cells From Patients With Polycystic Ovary... Reprod Sci. 2017 Jun;24(6):902-910. | |||||
| REF 2 | ClinicalTrials.gov (NCT04474470) A Study to Evaluate NT219 Alone and in Combination With ERBITUX (Cetuximab) in Adults With Advanced Solid Tumors and Head and Neck Cancer. U.S. National Institutes of Health. | |||||
| REF 3 | Structure and autoregulation of the insulin-like growth factor 1 receptor kinase. Nat Struct Biol. 2001 Dec;8(12):1058-63. | |||||
If You Find Any Error in Data or Bug in Web Service, Please Kindly Report It to Dr. Zhou and Dr. Zhang.

