Target Information
| Target General Information | Top | |||||
|---|---|---|---|---|---|---|
| Target ID |
T25464
(Former ID: TTDS00363)
|
|||||
| Target Name |
Calcineurin (PPP3CA)
|
|||||
| Synonyms |
Serine/threonine-protein phosphatase 2B catalytic subunit alpha isoform; Calmodulin-dependent calcineurin A subunit alpha isoform; CNA; CAM-PRP catalytic subunit; CALNA
Click to Show/Hide
|
|||||
| Gene Name |
PPP3CA
|
|||||
| Target Type |
Successful target
|
[1] | ||||
| Disease | [+] 4 Target-related Diseases | + | ||||
| 1 | Atopic eczema [ICD-11: EA80] | |||||
| 2 | Lupus erythematosus [ICD-11: 4A40] | |||||
| 3 | Psoriasis [ICD-11: EA90] | |||||
| 4 | Transplant rejection [ICD-11: NE84] | |||||
| Function |
Many of the substrates contain a PxIxIT motif and/or a LxVP motif. In response to increased Ca(2+) levels, dephosphorylates and activates phosphatase SSH1 which results in cofilin dephosphorylation. In response to increased Ca(2+) levels following mitochondrial depolarization, dephosphorylates DNM1L inducing DNM1L translocation to the mitochondrion. Dephosphorylates heat shock protein HSPB1. Dephosphorylates and activates transcription factor NFATC1. In response to increased Ca(2+) levels, regulates NFAT-mediated transcription probably by dephosphorylating NFAT and promoting its nuclear translocation. Dephosphorylates and inactivates transcription factor ELK1. Dephosphorylates DARPP32. May dephosphorylate CRTC2 at 'Ser-171' resulting in CRTC2 dissociation from 14-3-3 proteins. Calcium-dependent, calmodulin-stimulated protein phosphatase which plays an essential role in the transduction of intracellular Ca(2+)-mediated signals.
Click to Show/Hide
|
|||||
| BioChemical Class |
Phosphoric monoester hydrolase
|
|||||
| UniProt ID | ||||||
| EC Number |
EC 3.1.3.16
|
|||||
| Sequence |
MSEPKAIDPKLSTTDRVVKAVPFPPSHRLTAKEVFDNDGKPRVDILKAHLMKEGRLEESV
ALRIITEGASILRQEKNLLDIDAPVTVCGDIHGQFFDLMKLFEVGGSPANTRYLFLGDYV DRGYFSIECVLYLWALKILYPKTLFLLRGNHECRHLTEYFTFKQECKIKYSERVYDACMD AFDCLPLAALMNQQFLCVHGGLSPEINTLDDIRKLDRFKEPPAYGPMCDILWSDPLEDFG NEKTQEHFTHNTVRGCSYFYSYPAVCEFLQHNNLLSILRAHEAQDAGYRMYRKSQTTGFP SLITIFSAPNYLDVYNNKAAVLKYENNVMNIRQFNCSPHPYWLPNFMDVFTWSLPFVGEK VTEMLVNVLNICSDDELGSEEDGFDGATAAARKEVIRNKIRAIGKMARVFSVLREESESV LTLKGLTPTGMLPSGVLSGGKQTLQSATVEAIEADEAIKGFSPQHKITSFEEAKGLDRIN ERMPPRRDAMPSDANLNSINKALTSETNGTDSNGSNSSNIQ Click to Show/Hide
|
|||||
| 3D Structure | Click to Show 3D Structure of This Target | PDB | ||||
| HIT2.0 ID | T71DUF | |||||
| Drugs and Modes of Action | Top | |||||
|---|---|---|---|---|---|---|
| Approved Drug(s) | [+] 2 Approved Drugs | + | ||||
| 1 | Tacrolimus | Drug Info | Approved | Organ transplant rejection | [4], [5] | |
| 2 | Voclosporin | Drug Info | Approved | Lupus nephritis | [6] | |
| Mode of Action | [+] 1 Modes of Action | + | ||||
| Inhibitor | [+] 3 Inhibitor drugs | + | ||||
| 1 | Tacrolimus | Drug Info | [7] | |||
| 2 | Voclosporin | Drug Info | [6] | |||
| 3 | L-685,818 | Drug Info | [8] | |||
| Cell-based Target Expression Variations | Top | |||||
|---|---|---|---|---|---|---|
| Cell-based Target Expression Variations | ||||||
| Different Human System Profiles of Target | Top |
|---|---|
|
Human Similarity Proteins
of target is determined by comparing the sequence similarity of all human proteins with the target based on BLAST. The similarity proteins for a target are defined as the proteins with E-value < 0.005 and outside the protein families of the target.
A target that has fewer human similarity proteins outside its family is commonly regarded to possess a greater capacity to avoid undesired interactions and thus increase the possibility of finding successful drugs
(Brief Bioinform, 21: 649-662, 2020).
Human Tissue Distribution
of target is determined from a proteomics study that quantified more than 12,000 genes across 32 normal human tissues. Tissue Specificity (TS) score was used to define the enrichment of target across tissues.
The distribution of targets among different tissues or organs need to be taken into consideration when assessing the target druggability, as it is generally accepted that the wider the target distribution, the greater the concern over potential adverse effects
(Nat Rev Drug Discov, 20: 64-81, 2021).
Human Pathway Affiliation
of target is determined by the life-essential pathways provided on KEGG database. The target-affiliated pathways were defined based on the following two criteria (a) the pathways of the studied target should be life-essential for both healthy individuals and patients, and (b) the studied target should occupy an upstream position in the pathways and therefore had the ability to regulate biological function.
Targets involved in a fewer pathways have greater likelihood to be successfully developed, while those associated with more human pathways increase the chance of undesirable interferences with other human processes
(Pharmacol Rev, 58: 259-279, 2006).
Biological Network Descriptors
of target is determined based on a human protein-protein interactions (PPI) network consisting of 9,309 proteins and 52,713 PPIs, which were with a high confidence score of ≥ 0.95 collected from STRING database.
The network properties of targets based on protein-protein interactions (PPIs) have been widely adopted for the assessment of target’s druggability. Proteins with high node degree tend to have a high impact on network function through multiple interactions, while proteins with high betweenness centrality are regarded to be central for communication in interaction networks and regulate the flow of signaling information
(Front Pharmacol, 9, 1245, 2018;
Curr Opin Struct Biol. 44:134-142, 2017).
Human Similarity Proteins
Human Tissue Distribution
Human Pathway Affiliation
Biological Network Descriptors
|
|
|
There is no similarity protein (E value < 0.005) for this target
|
|
Note:
If a protein has TS (tissue specficity) scores at least in one tissue >= 2.5, this protein is called tissue-enriched (including tissue-enriched-but-not-specific and tissue-specific). In the plots, the vertical lines are at thresholds 2.5 and 4.
|
| KEGG Pathway | Pathway ID | Affiliated Target | Pathway Map |
|---|---|---|---|
| MAPK signaling pathway | hsa04010 | Affiliated Target |
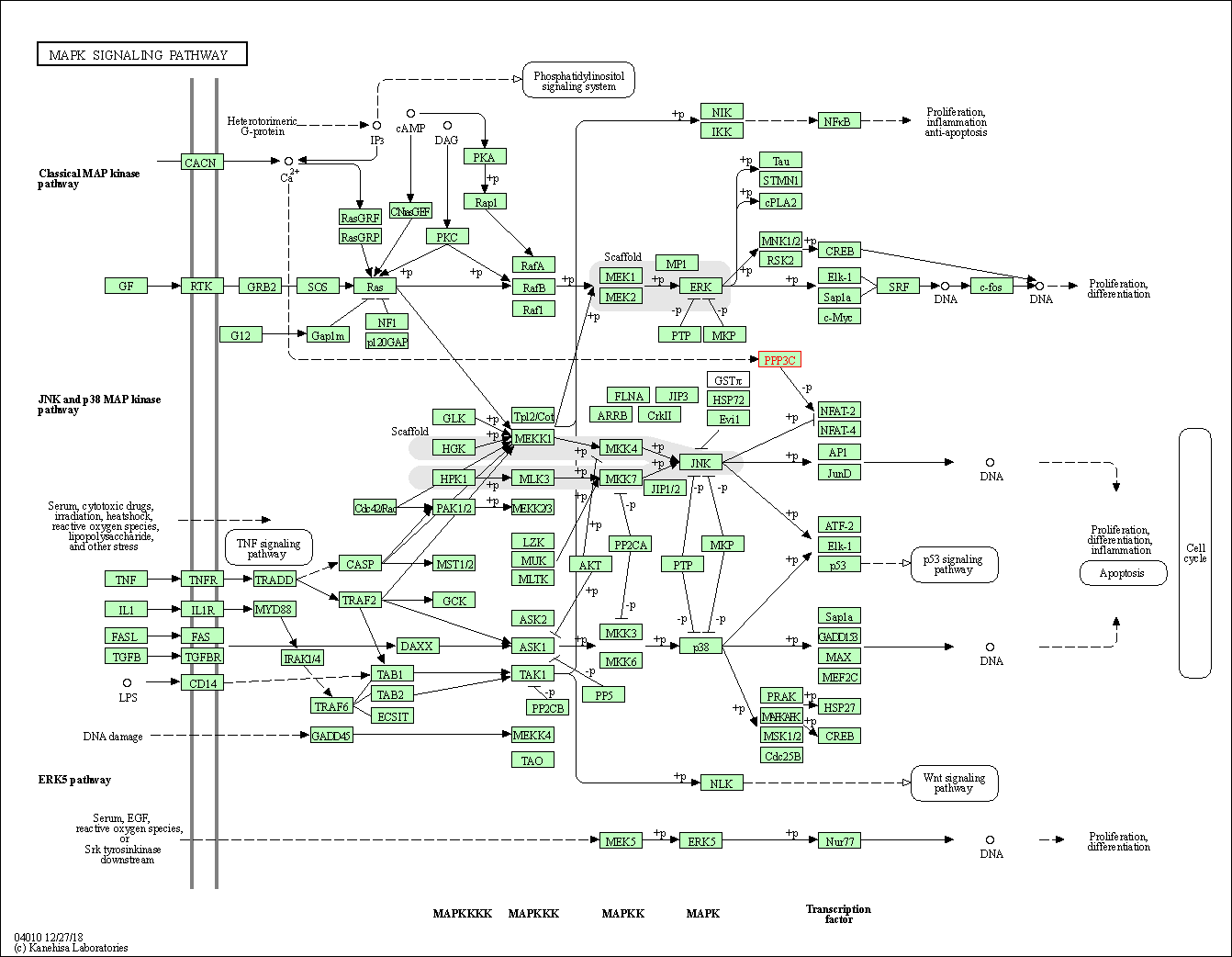
|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Calcium signaling pathway | hsa04020 | Affiliated Target |
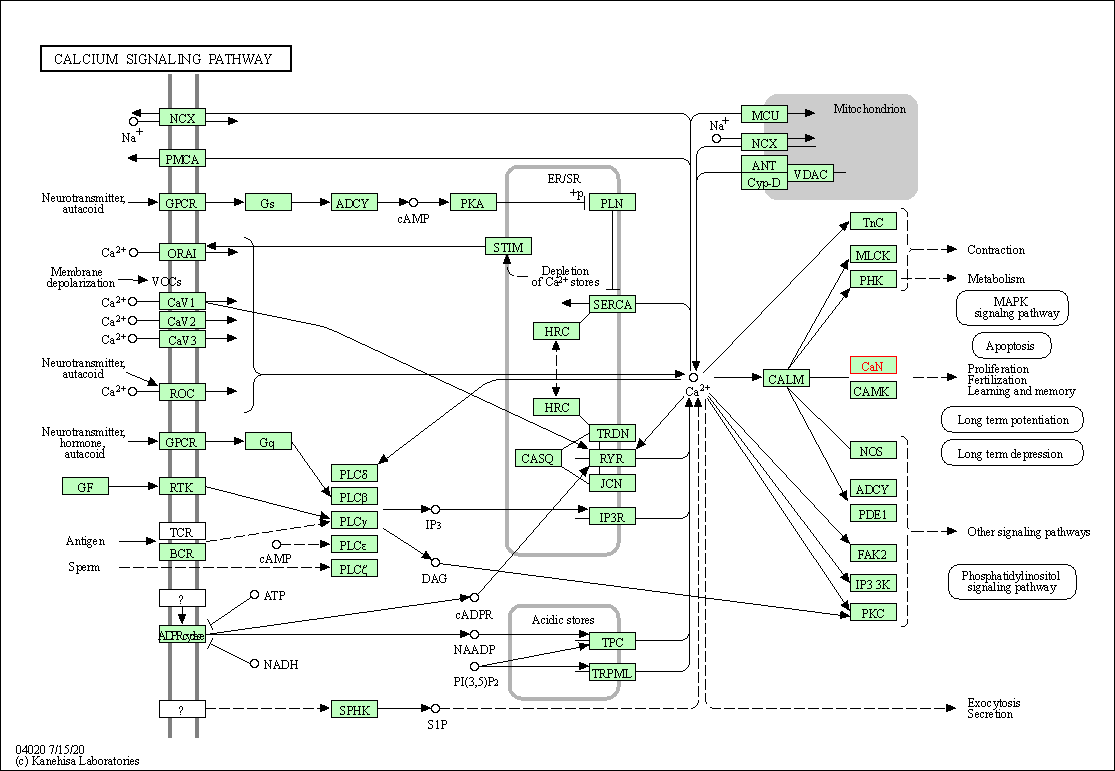
|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| cGMP-PKG signaling pathway | hsa04022 | Affiliated Target |
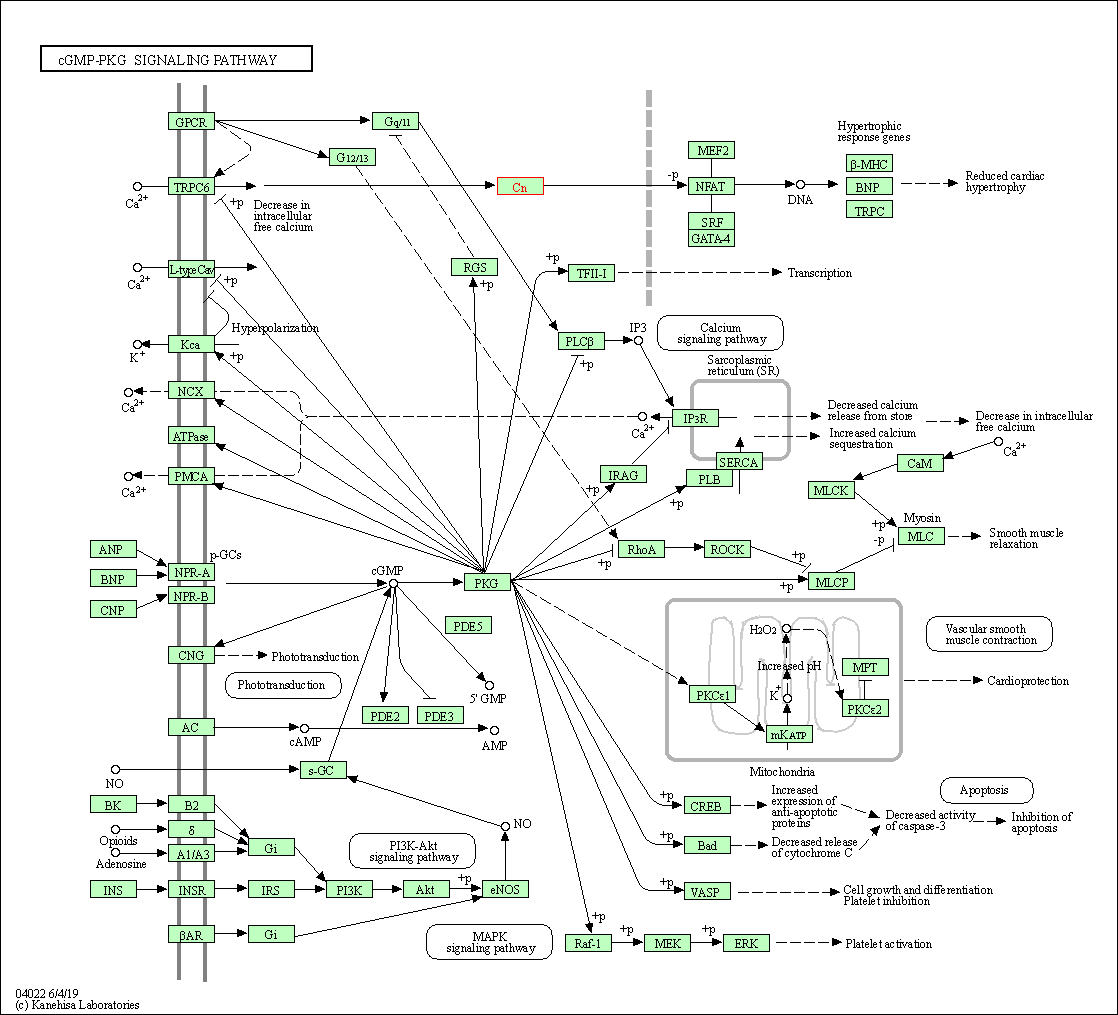
|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Oocyte meiosis | hsa04114 | Affiliated Target |
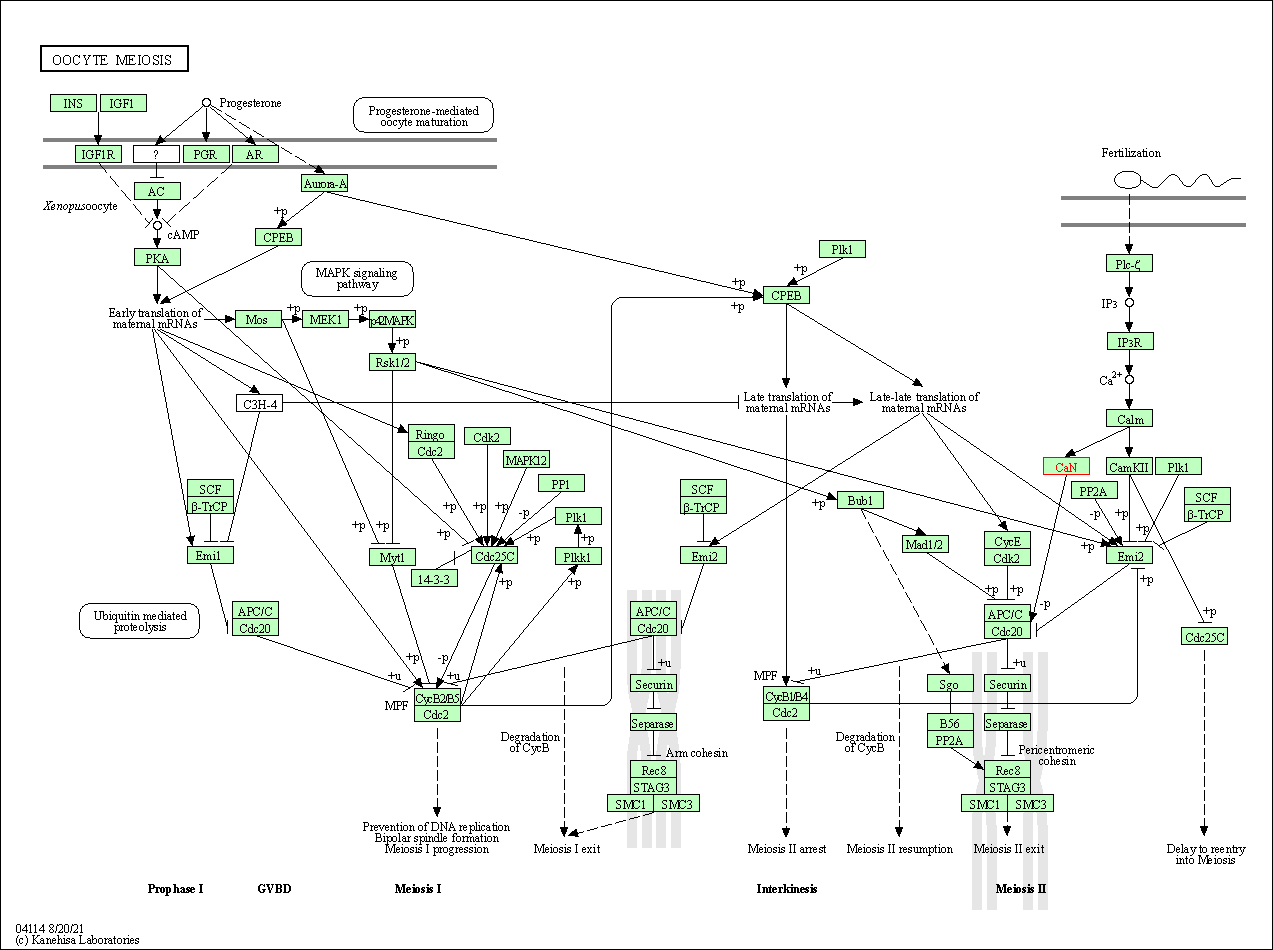
|
| Class: Cellular Processes => Cell growth and death | Pathway Hierarchy | ||
| Cellular senescence | hsa04218 | Affiliated Target |
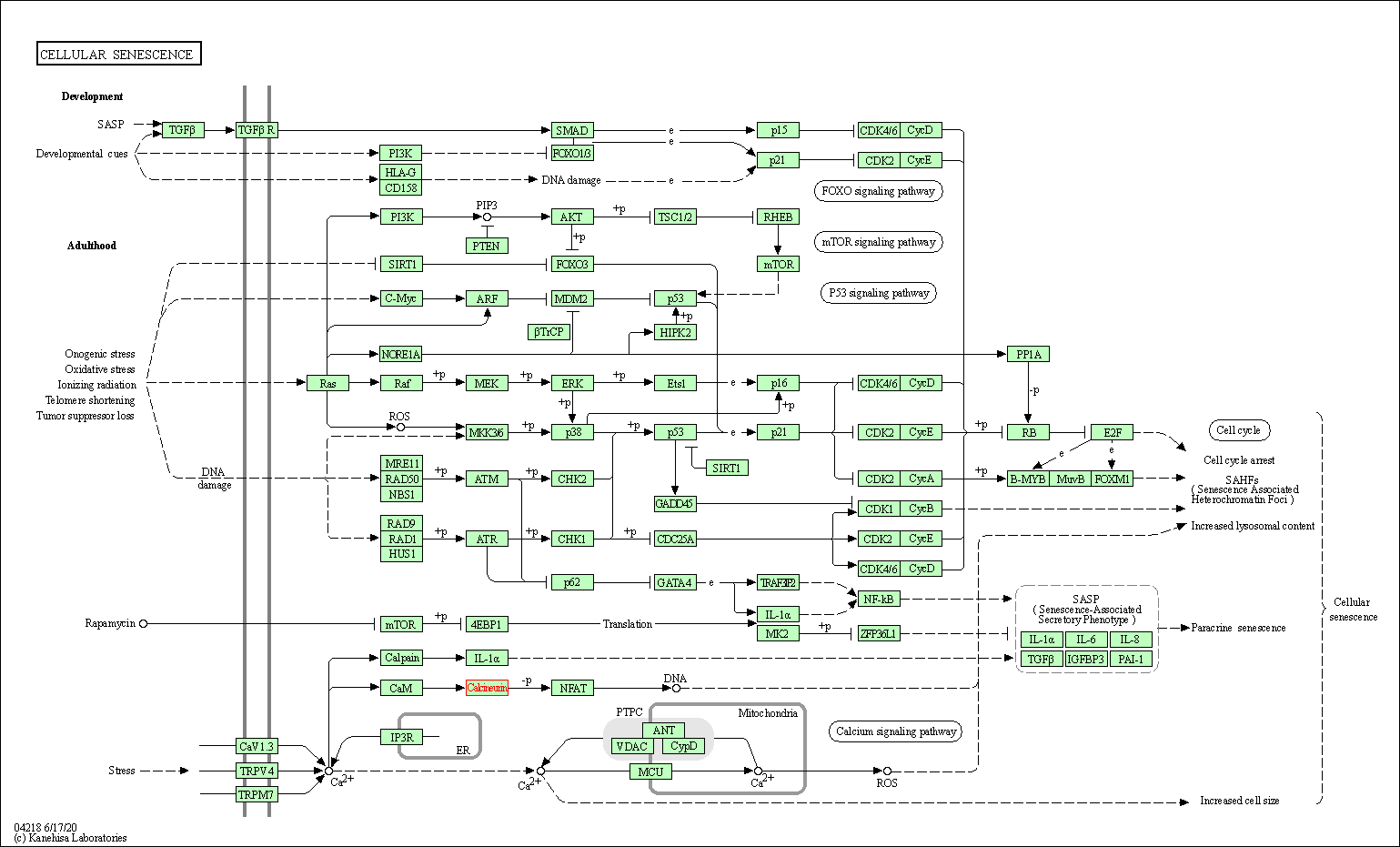
|
| Class: Cellular Processes => Cell growth and death | Pathway Hierarchy | ||
| Wnt signaling pathway | hsa04310 | Affiliated Target |
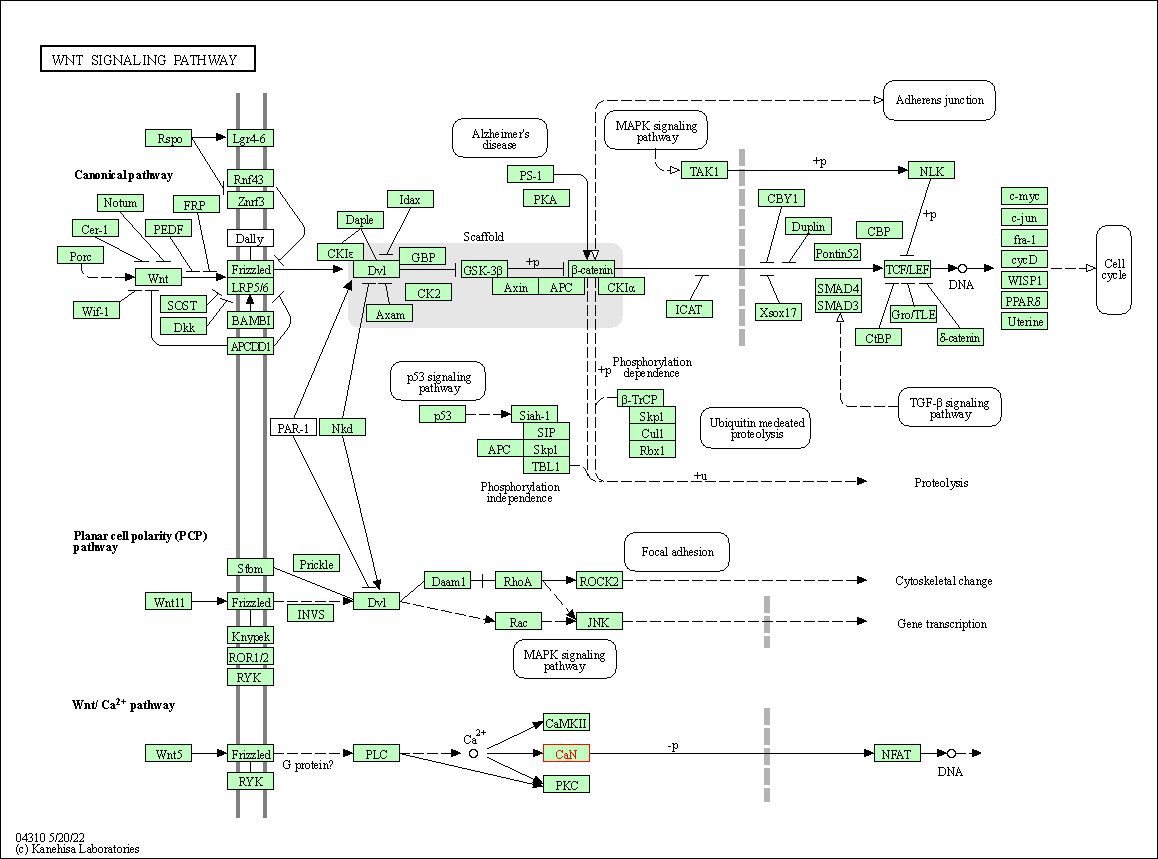
|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Axon guidance | hsa04360 | Affiliated Target |
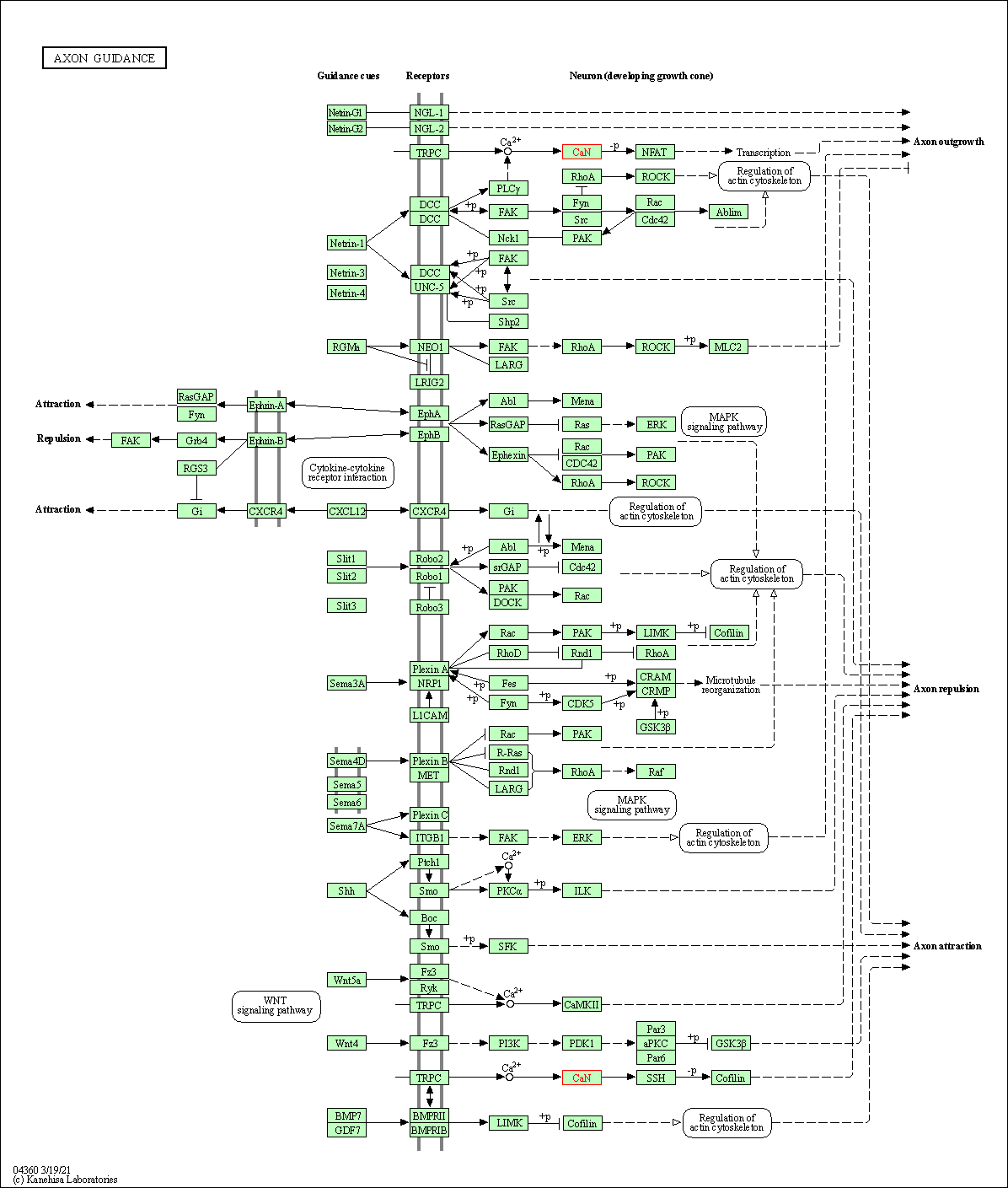
|
| Class: Organismal Systems => Development and regeneration | Pathway Hierarchy | ||
| VEGF signaling pathway | hsa04370 | Affiliated Target |
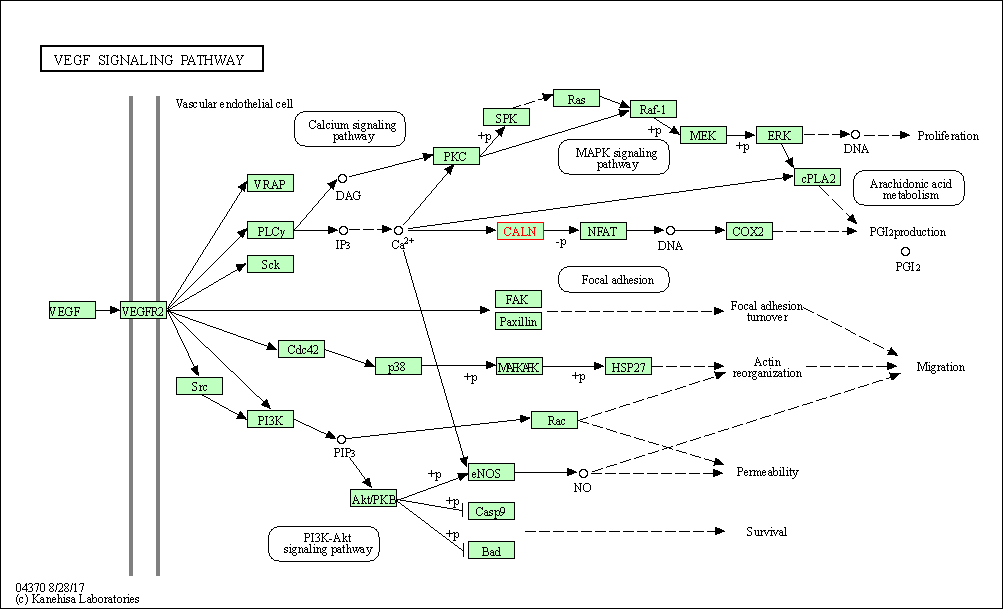
|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Osteoclast differentiation | hsa04380 | Affiliated Target |
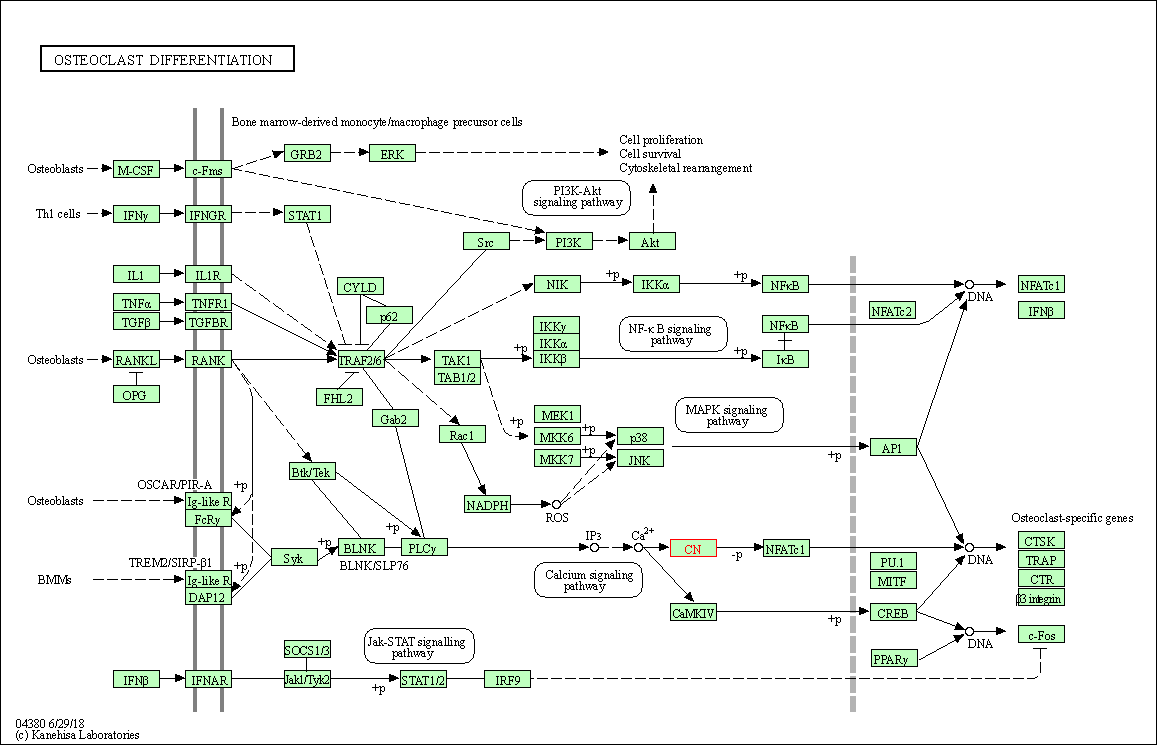
|
| Class: Organismal Systems => Development and regeneration | Pathway Hierarchy | ||
| C-type lectin receptor signaling pathway | hsa04625 | Affiliated Target |
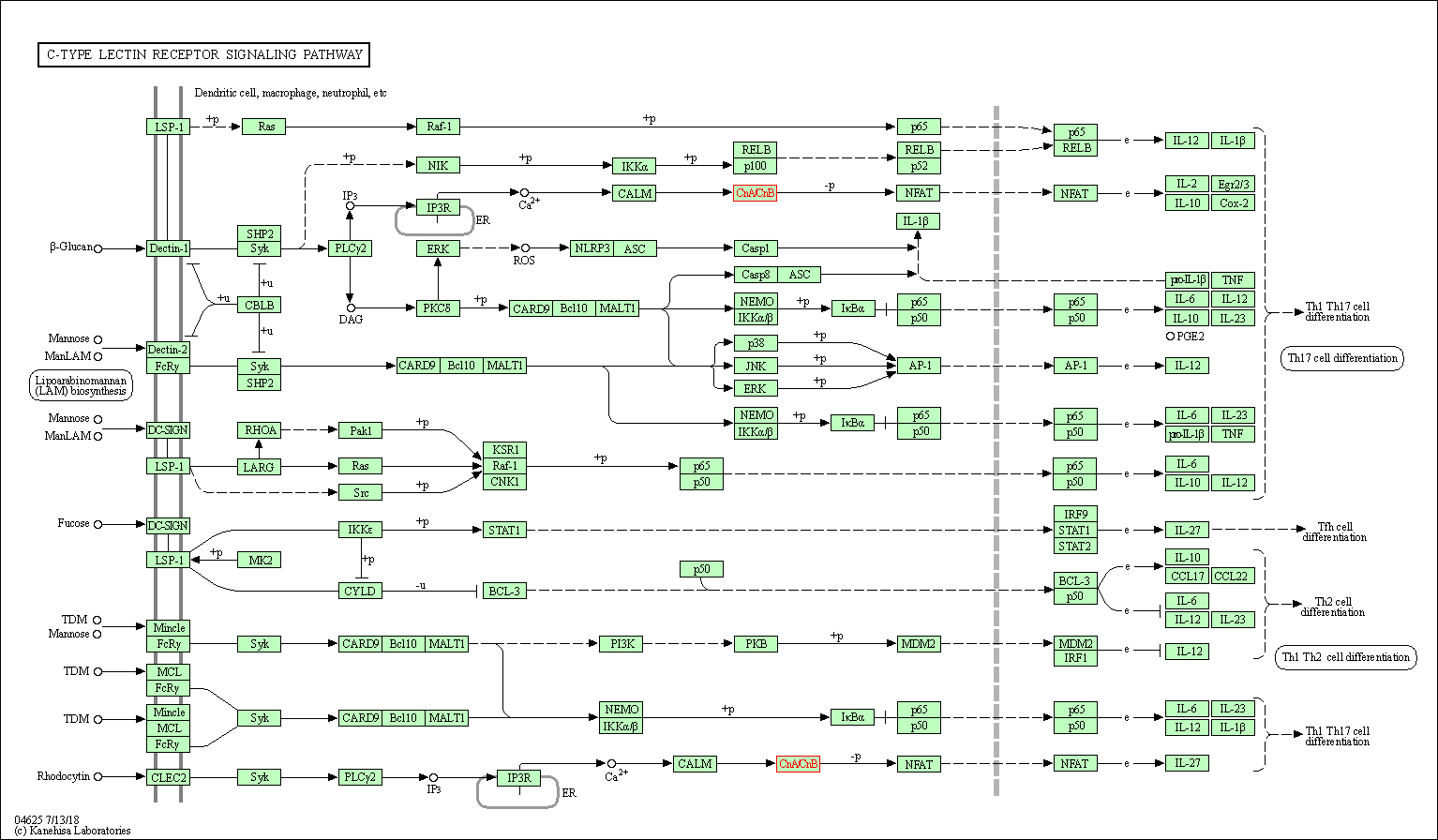
|
| Class: Organismal Systems => Immune system | Pathway Hierarchy | ||
| Natural killer cell mediated cytotoxicity | hsa04650 | Affiliated Target |
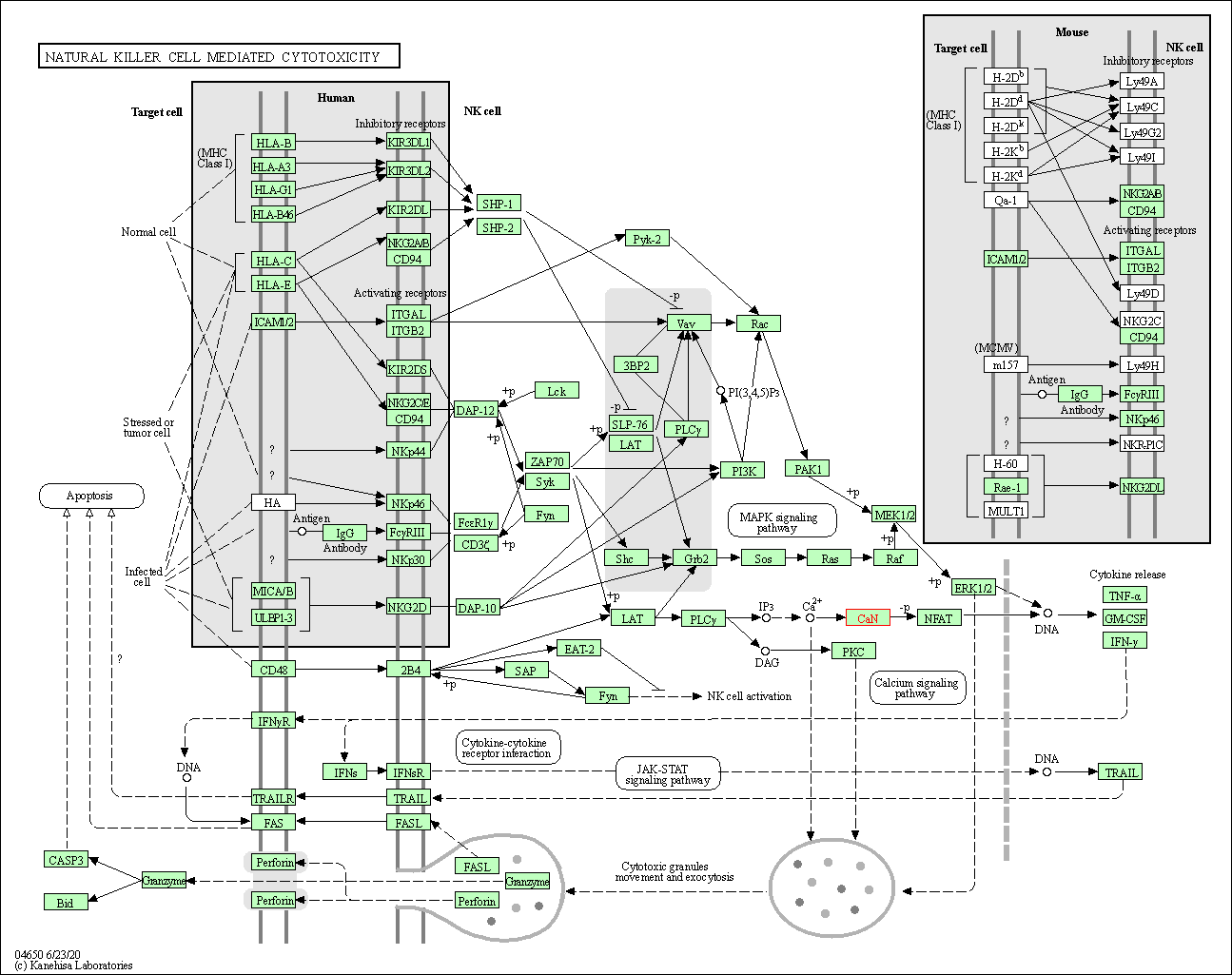
|
| Class: Organismal Systems => Immune system | Pathway Hierarchy | ||
| Th1 and Th2 cell differentiation | hsa04658 | Affiliated Target |
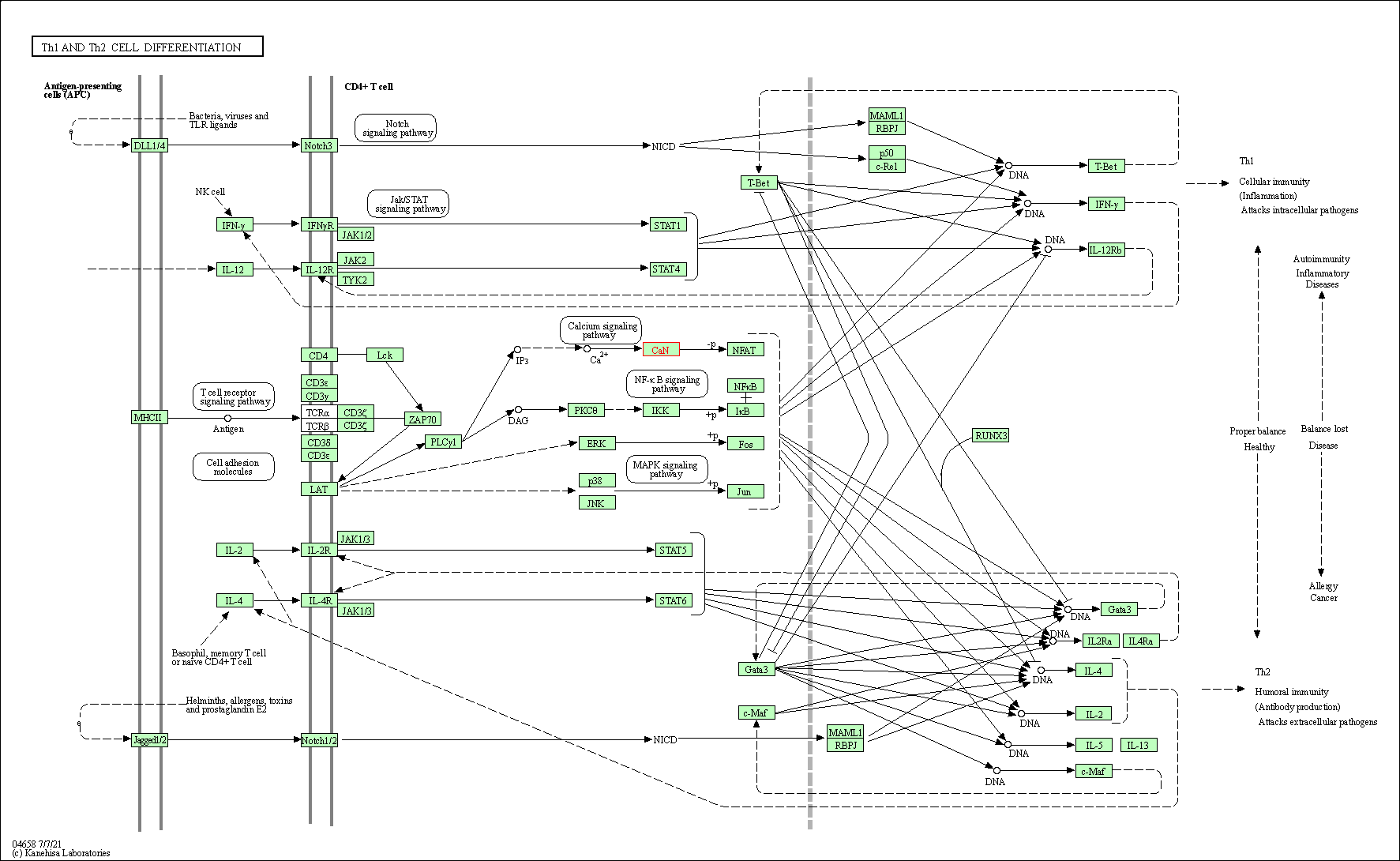
|
| Class: Organismal Systems => Immune system | Pathway Hierarchy | ||
| Th17 cell differentiation | hsa04659 | Affiliated Target |
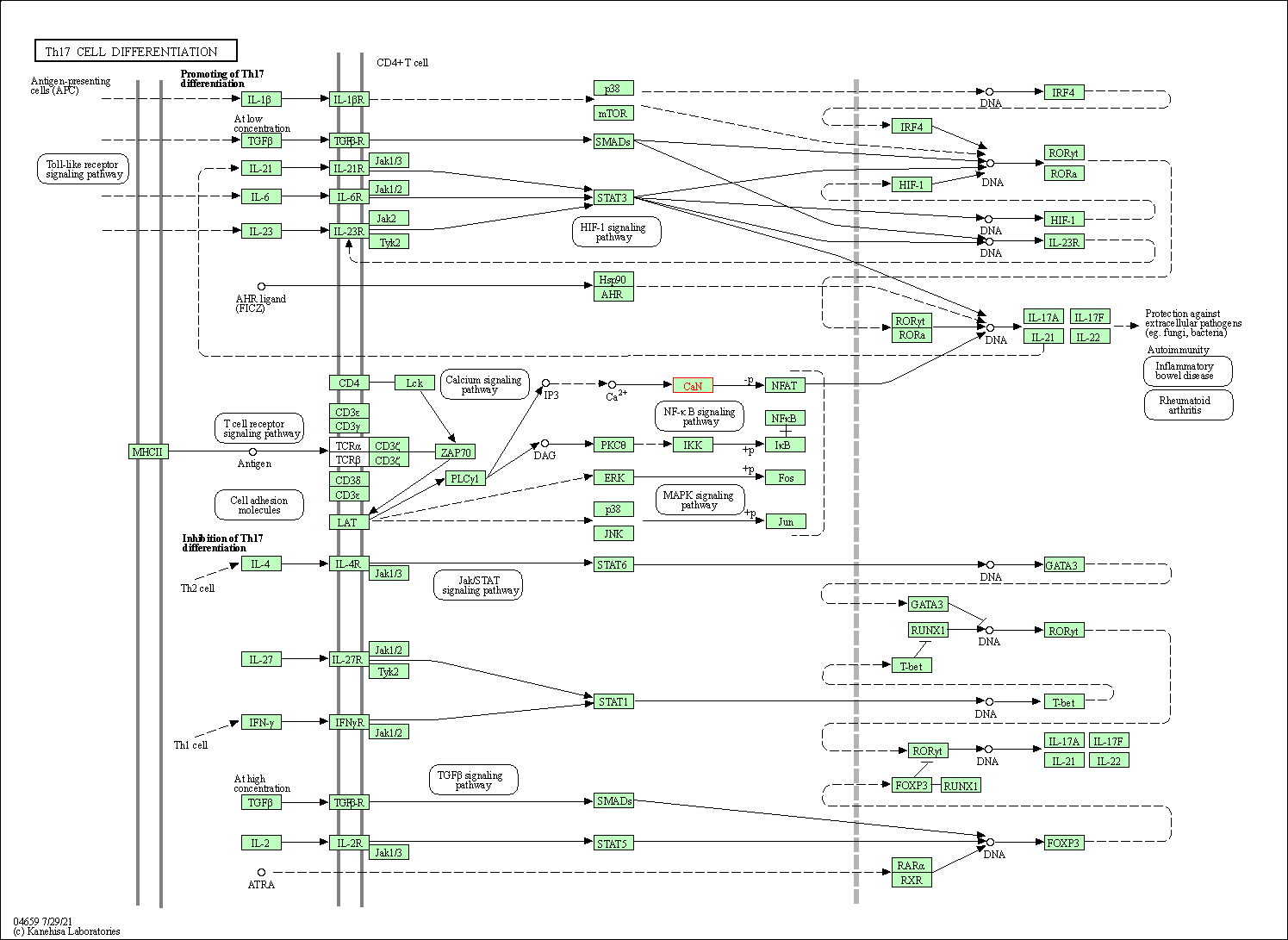
|
| Class: Organismal Systems => Immune system | Pathway Hierarchy | ||
| T cell receptor signaling pathway | hsa04660 | Affiliated Target |
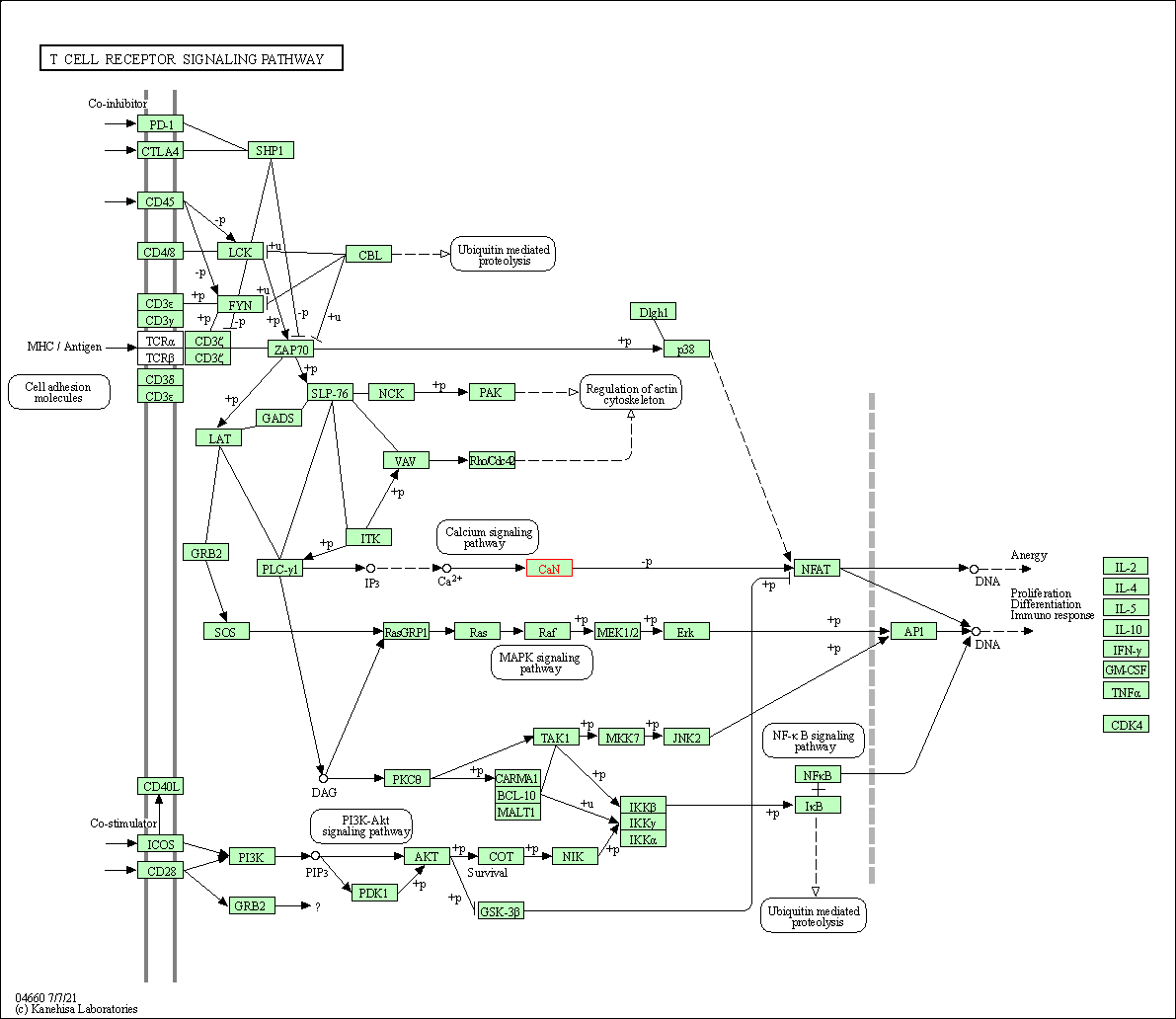
|
| Class: Organismal Systems => Immune system | Pathway Hierarchy | ||
| B cell receptor signaling pathway | hsa04662 | Affiliated Target |
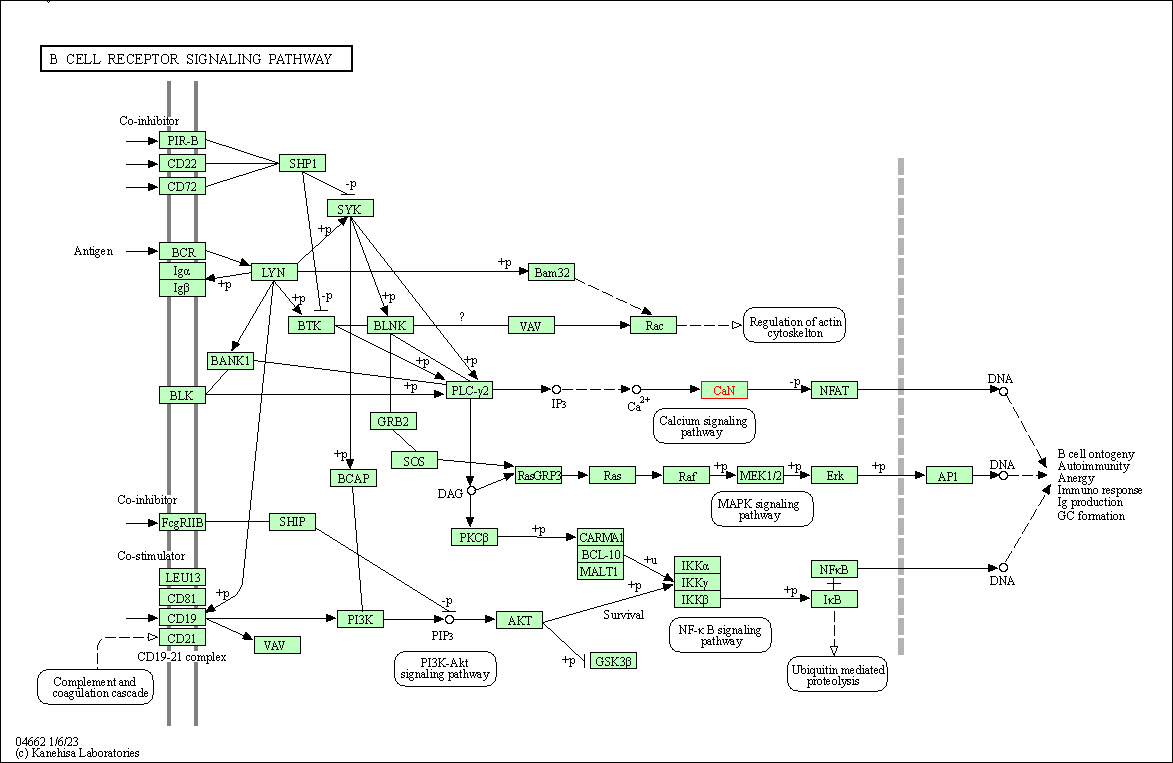
|
| Class: Organismal Systems => Immune system | Pathway Hierarchy | ||
| Long-term potentiation | hsa04720 | Affiliated Target |
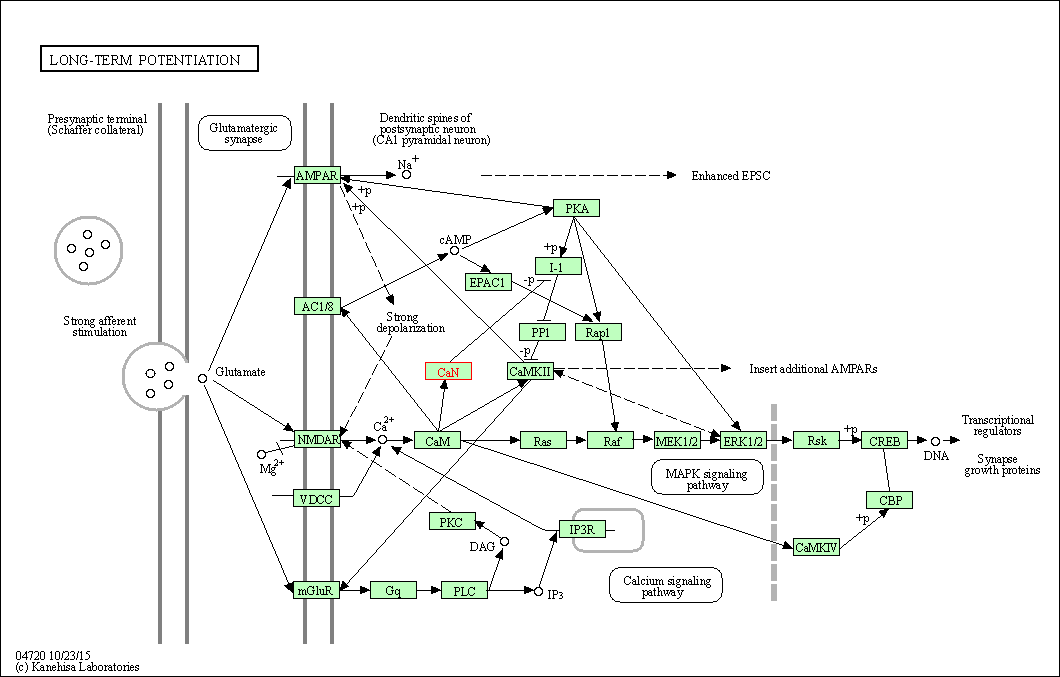
|
| Class: Organismal Systems => Nervous system | Pathway Hierarchy | ||
| Glutamatergic synapse | hsa04724 | Affiliated Target |
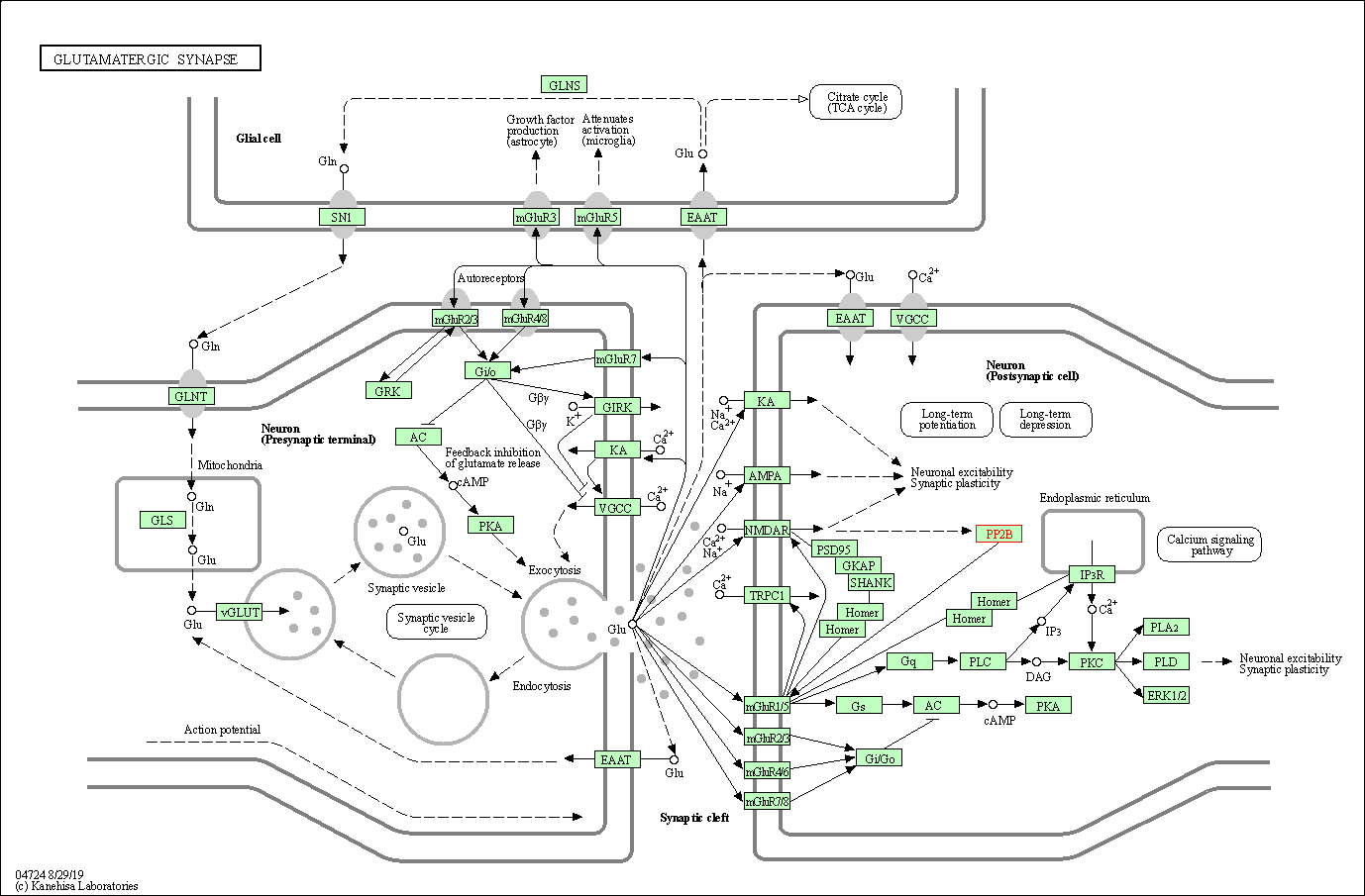
|
| Class: Organismal Systems => Nervous system | Pathway Hierarchy | ||
| Dopaminergic synapse | hsa04728 | Affiliated Target |
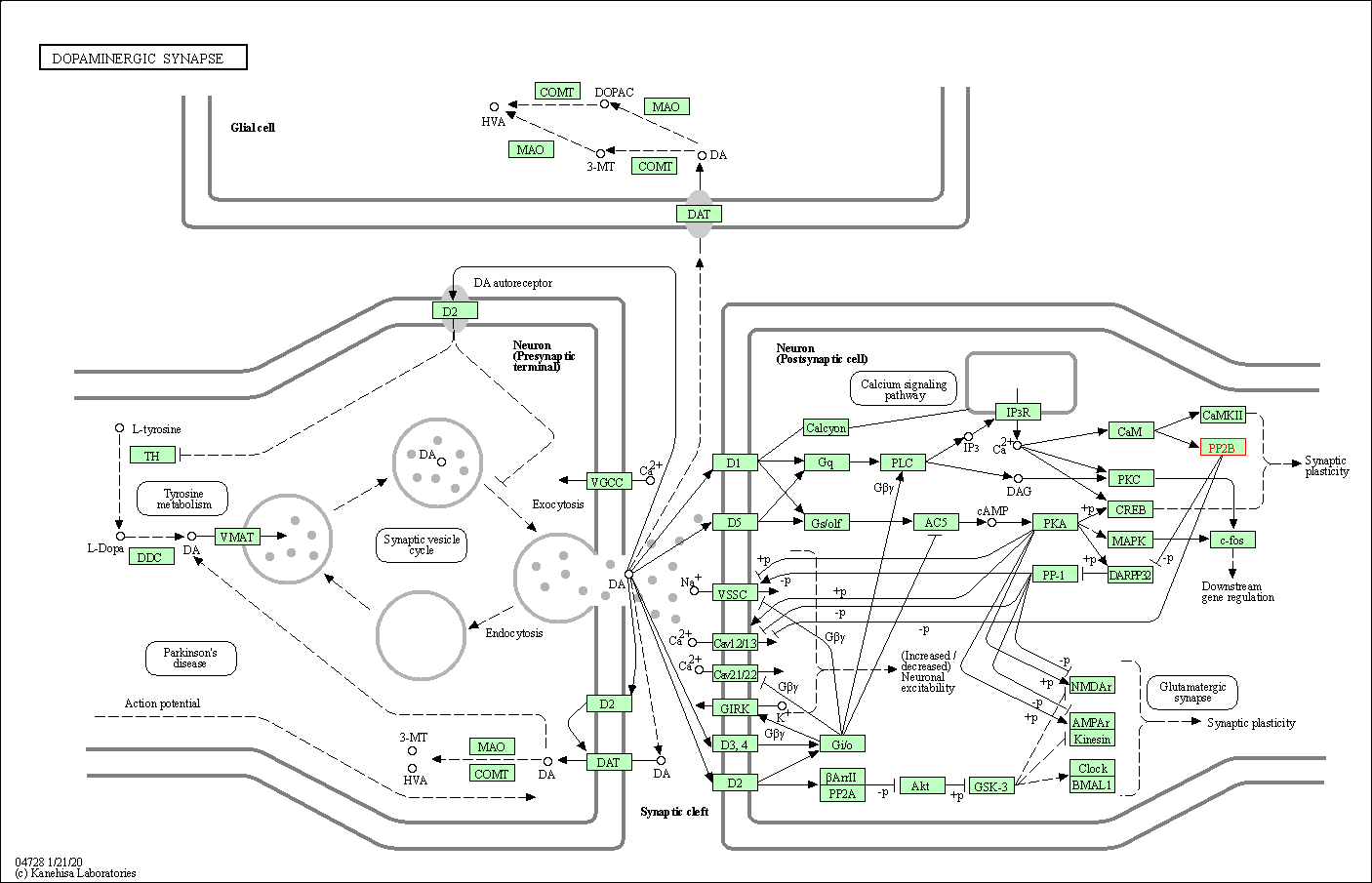
|
| Class: Organismal Systems => Nervous system | Pathway Hierarchy | ||
| Oxytocin signaling pathway | hsa04921 | Affiliated Target |
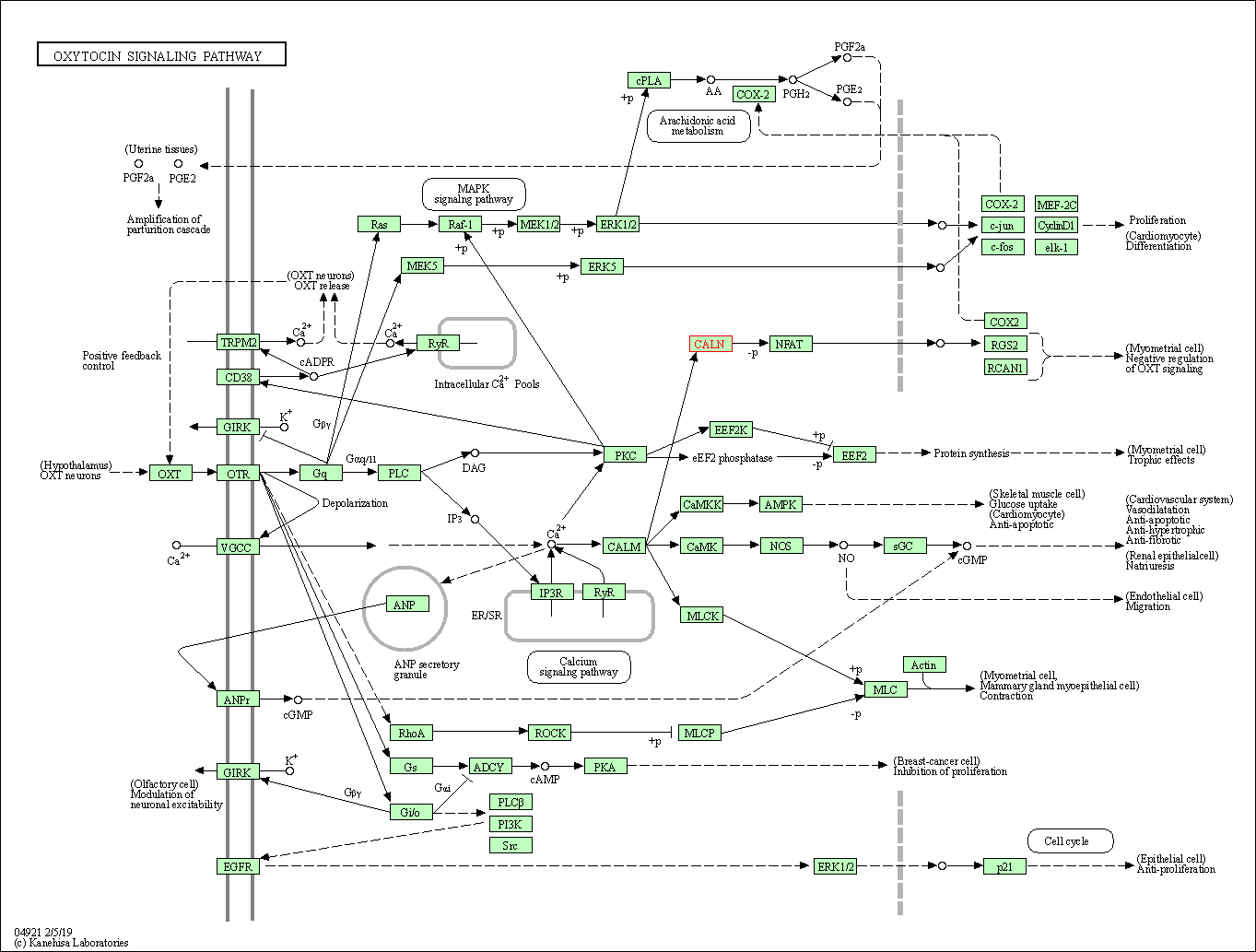
|
| Class: Organismal Systems => Endocrine system | Pathway Hierarchy | ||
| Glucagon signaling pathway | hsa04922 | Affiliated Target |
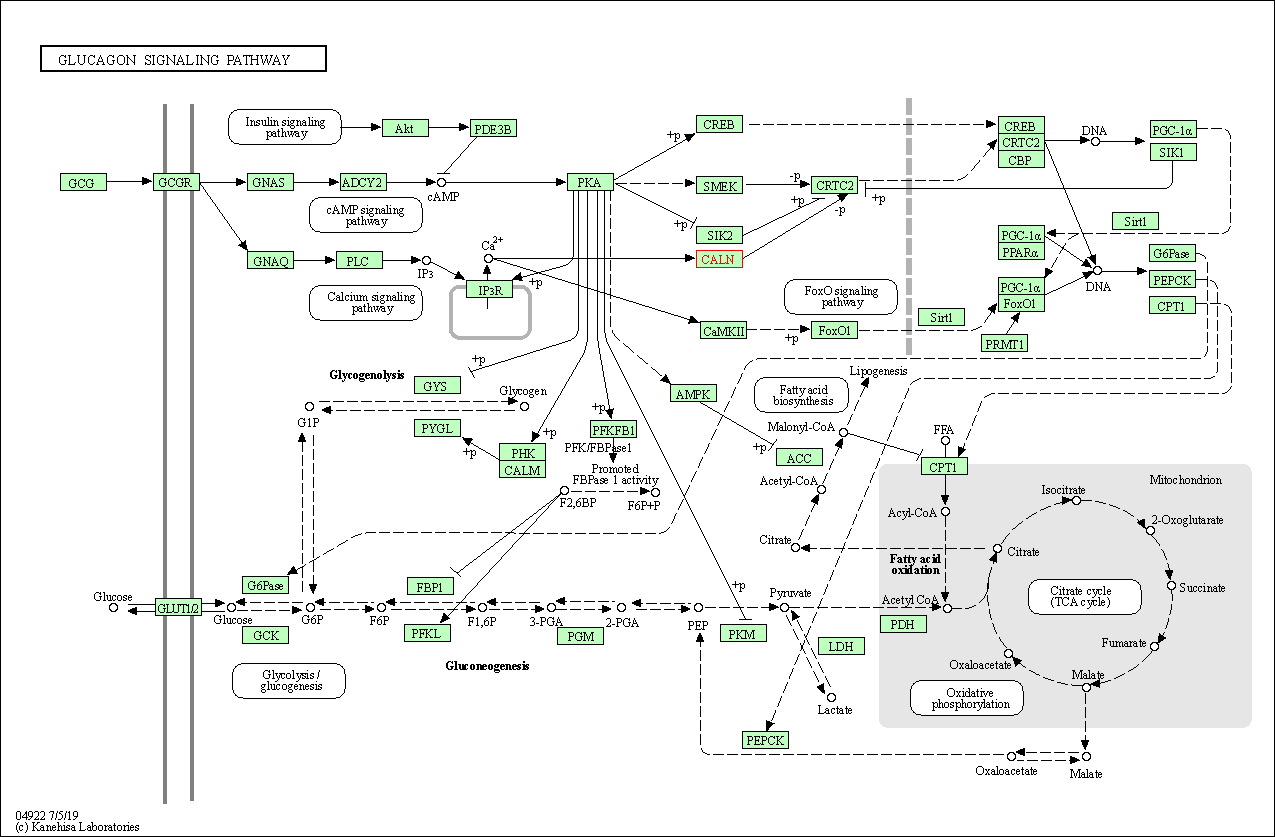
|
| Class: Organismal Systems => Endocrine system | Pathway Hierarchy | ||
| Renin secretion | hsa04924 | Affiliated Target |
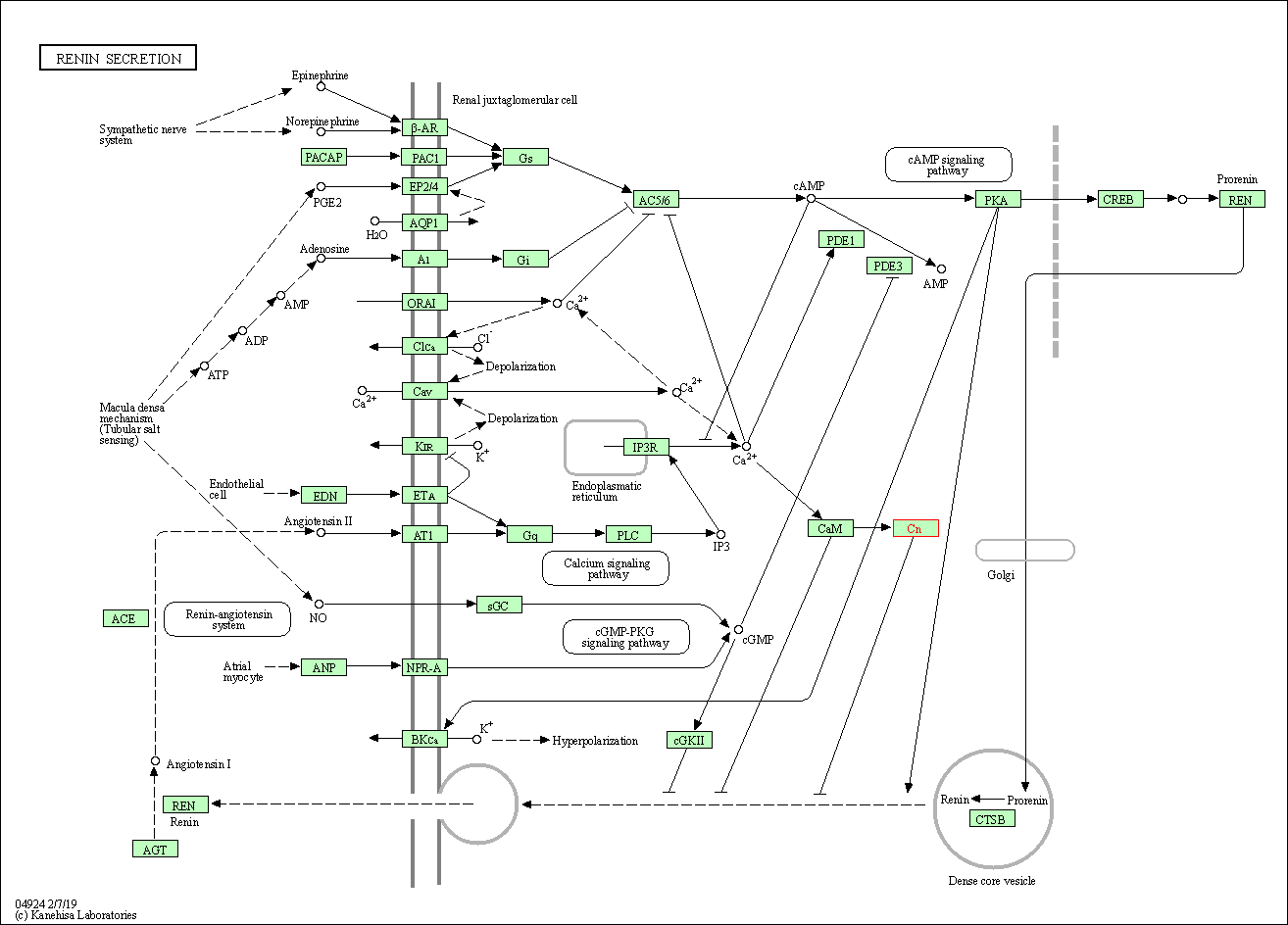
|
| Class: Organismal Systems => Endocrine system | Pathway Hierarchy | ||
| Click to Show/Hide the Information of Affiliated Human Pathways | |||
| Degree | 10 | Degree centrality | 1.07E-03 | Betweenness centrality | 6.01E-04 |
|---|---|---|---|---|---|
| Closeness centrality | 2.06E-01 | Radiality | 1.36E+01 | Clustering coefficient | 8.89E-02 |
| Neighborhood connectivity | 8.60E+00 | Topological coefficient | 1.22E-01 | Eccentricity | 12 |
| Download | Click to Download the Full PPI Network of This Target | ||||
| Chemical Structure based Activity Landscape of Target | Top |
|---|---|
| Drug Property Profile of Target | Top | |
|---|---|---|
| (1) Molecular Weight (mw) based Drug Clustering | (2) Octanol/Water Partition Coefficient (xlogp) based Drug Clustering | |
|
|
||
| (3) Hydrogen Bond Donor Count (hbonddonor) based Drug Clustering | (4) Hydrogen Bond Acceptor Count (hbondacc) based Drug Clustering | |
|
|
||
| (5) Rotatable Bond Count (rotbonds) based Drug Clustering | (6) Topological Polar Surface Area (polararea) based Drug Clustering | |
|
|
||
| "RO5" indicates the cutoff set by lipinski's rule of five; "D123AB" colored in GREEN denotes the no violation of any cutoff in lipinski's rule of five; "D123AB" colored in PURPLE refers to the violation of only one cutoff in lipinski's rule of five; "D123AB" colored in BLACK represents the violation of more than one cutoffs in lipinski's rule of five | ||
| Co-Targets | Top | |||||
|---|---|---|---|---|---|---|
| Co-Targets | ||||||
| Target Poor or Non Binders | Top | |||||
|---|---|---|---|---|---|---|
| Target Poor or Non Binders | ||||||
| Target Regulators | Top | |||||
|---|---|---|---|---|---|---|
| Target-regulating microRNAs | ||||||
| Target-interacting Proteins | ||||||
| Target Profiles in Patients | Top | |||||
|---|---|---|---|---|---|---|
| Target Expression Profile (TEP) | ||||||
| Target-Related Models and Studies | Top | |||||
|---|---|---|---|---|---|---|
| Target Validation | ||||||
| References | Top | |||||
|---|---|---|---|---|---|---|
| REF 1 | Cyclosporine and tacrolimus for the treatment of rheumatoid arthritis. Curr Opin Rheumatol. 2007 May;19(3):238-45. | |||||
| REF 2 | URL: http://www.guidetopharmacology.org Nucleic Acids Res. 2015 Oct 12. pii: gkv1037. The IUPHAR/BPS Guide to PHARMACOLOGY in 2016: towards curated quantitative interactions between 1300 protein targets and 6000 ligands. (Ligand id: 6783). | |||||
| REF 3 | Emerging drugs for moderate-to-severe psoriasis. Expert Opin Emerg Drugs. 2005 Feb;10(1):35-52. | |||||
| REF 4 | URL: http://www.guidetopharmacology.org Nucleic Acids Res. 2015 Oct 12. pii: gkv1037. The IUPHAR/BPS Guide to PHARMACOLOGY in 2016: towards curated quantitative interactions between 1300 protein targets and 6000 ligands. (Ligand id: 6784). | |||||
| REF 5 | New developments in immunosuppressive therapy for heart transplantation. Expert Opin Emerg Drugs. 2009 Mar;14(1):1-21. | |||||
| REF 6 | Drugs@FDA. U.S. Food and Drug Administration. U.S. Department of Health Human Services. 2021 | |||||
| REF 7 | Emerging drugs for ocular allergy. Expert Opin Emerg Drugs. 2005 Aug;10(3):505-20. | |||||
| REF 8 | Synergistic antifungal activities of bafilomycin A(1), fluconazole, and the pneumocandin MK-0991/caspofungin acetate (L-743,873) with calcineurin inhibitors FK506 and L-685,818 against Cryptococcus neoformans. Antimicrob Agents Chemother. 2000 Mar;44(3):739-46. | |||||
If You Find Any Error in Data or Bug in Web Service, Please Kindly Report It to Dr. Zhou and Dr. Zhang.

